Whole transcriptome single cell analysis for FFPE tissues
/in News, Parse/by Harshita SharmaUnlock with quick sign up!
Key Takeaways
- Whole transcriptome single cell profiling from archival FFPE breast cancer samples resolves tumor, stromal, and immune compartments in a single experiment
- Distinct epithelial programs are identified across clinically relevant subtypes, including ER+, ER/PR+, HER2+, and TNBC
- Subtype- and proliferation-associated lncRNA expression patterns are captured, highlighting the value of unbiased RNA profiling in FFPE samples
Experimental Design:
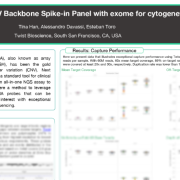
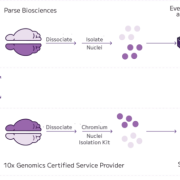
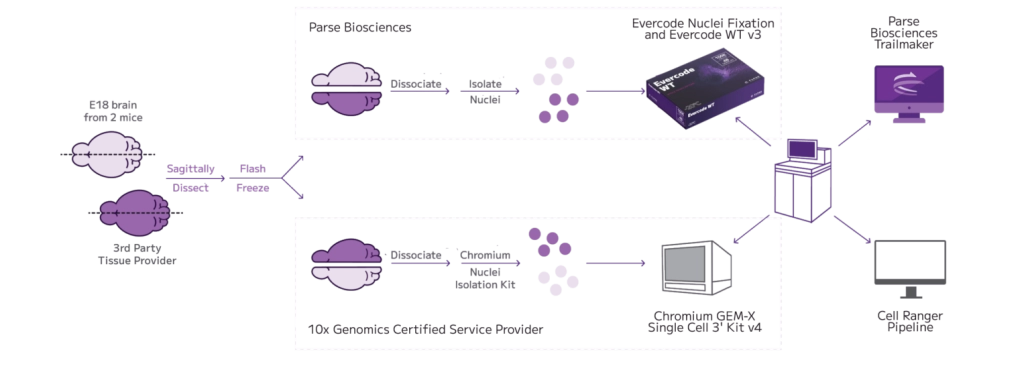
Formalin-fixed, paraffin-embedded (FFPE) tissue represents a vast source of clinically annotated samples, but has been difficult to use for whole transcriptome single cell analysis. In this dataset, nuclei isolated from 4 archived breast cancer FFPE samples were profiled using Evercode WT FFPE’s reverse transcription–based workflow designed to capture whole transcriptome expression from degraded RNA, profiling over 100,000 nuclei.
These results demonstrate that FFPE samples enable whole transcriptome profiling that captures meaningful cell types and tumor subtype biology across multiple donors while preserving cellular heterogeneity.
Results:
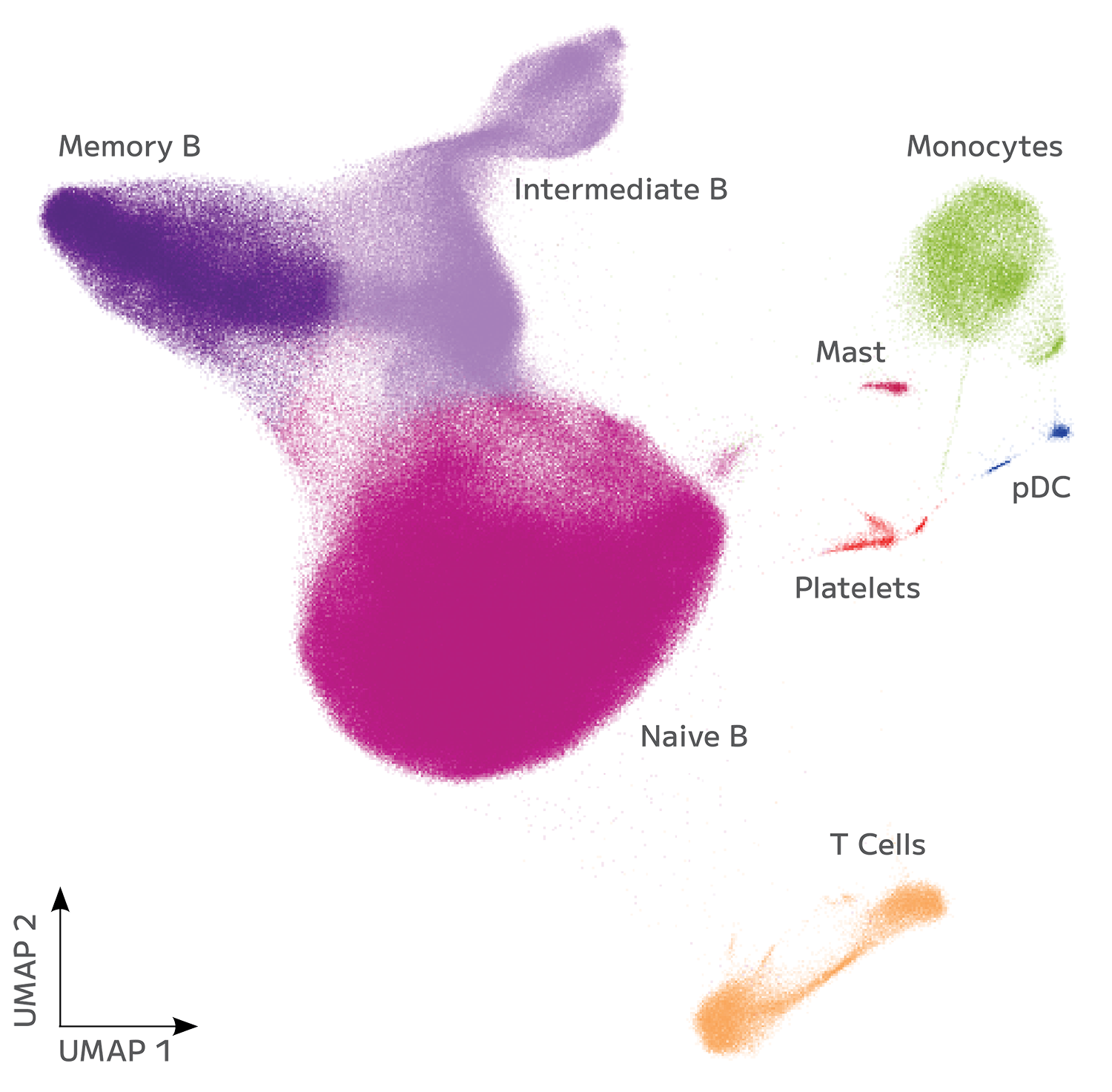
Whole transcriptome profiling resolved epithelial tumor populations alongside stromal and immune compartments, including CAFs, endothelial cells, VSMCs, myeloid cells, pDCs, B cells, NKT cells, and mast cells.
Cells cluster strongly by donor, with clear differences in gene expression programs across samples. Subtype-specific expression patterns distinguish ER+, ER/PR+, HER2+, and TNBC tumors, with TNBC remaining particularly distinct after integration.
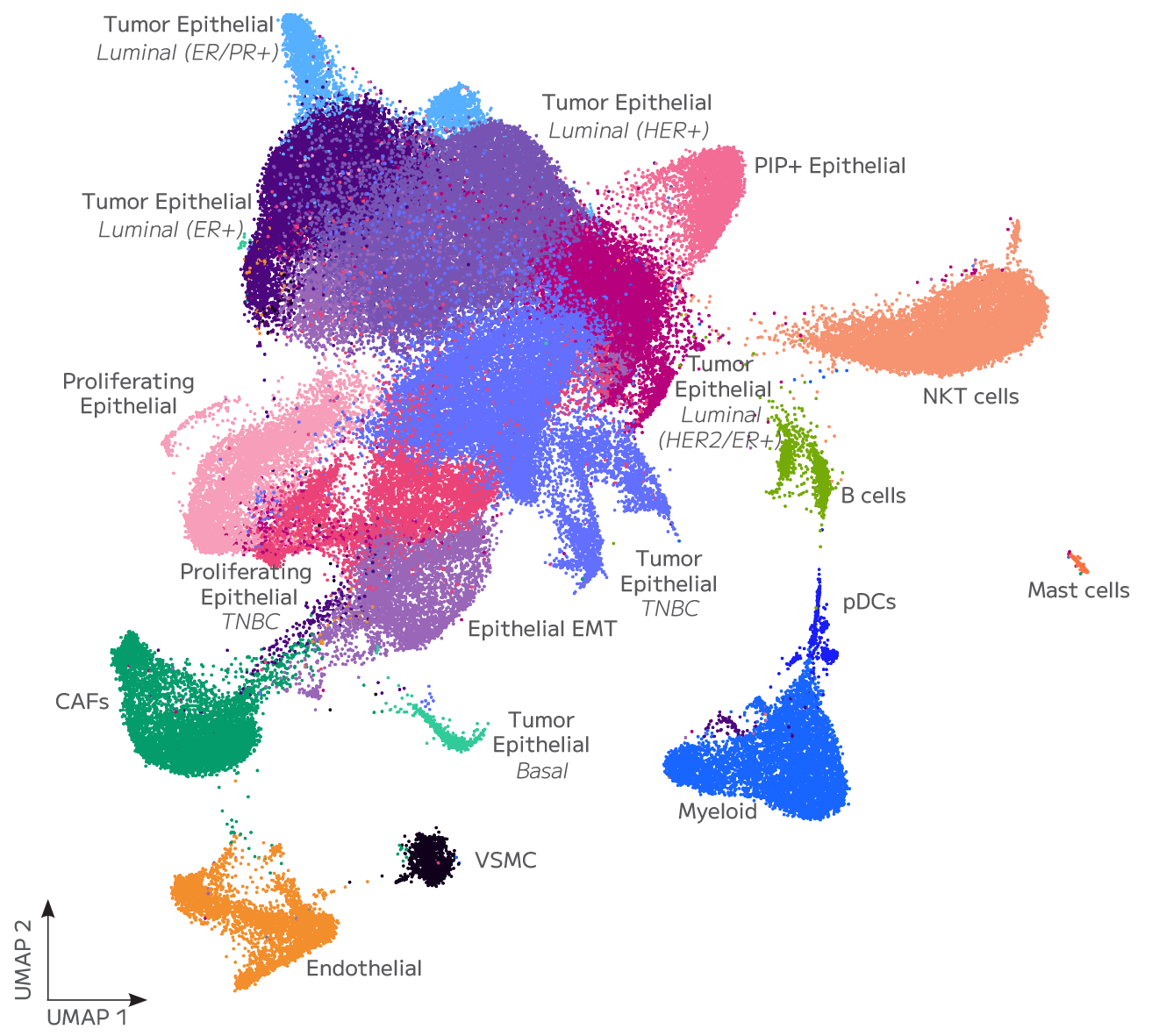
Figure 1: UMAP of human breast cancer FFPE nuclei. Tumor and proliferative epithelial states are resolved together with stromal and immune populations from FFPE samples.
Subtype- and state-associated lncRNA expression
Whole transcriptome profiling enables detection of biologically relevant lncRNAs across breast tumor cell states, revealing patterns linked to both tumor subtype and functional cell state.
LINC00993, a lncRNA associated with tumor-suppressive activity in breast cancer, is enriched in luminal epithelial populations. In contrast, the oncogenic lncRNA TUG1 shows higher expression in TNBC and proliferating epithelial states. These patterns are consistent with known subtype-associated biology and highlight how lncRNA expression reflects underlying tumor programs.
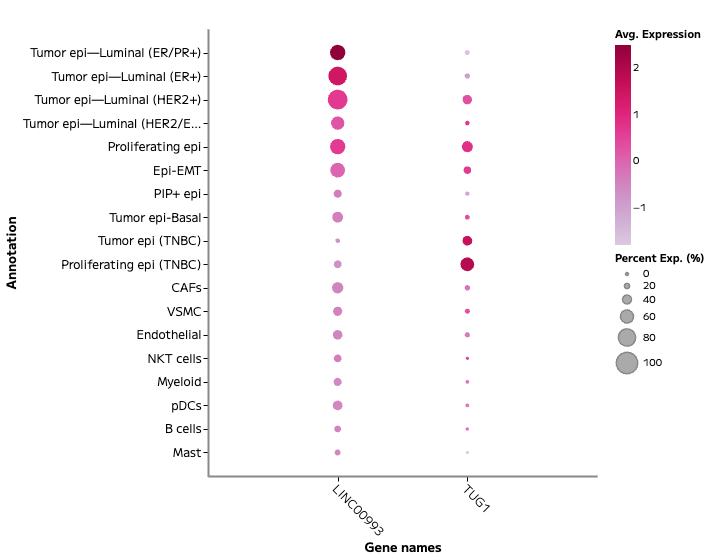
Figure 3: Proliferation-associated lncRNA expression.
Expression of TUG1 and additional lncRNAs across proliferating epithelial populations marked by Ki67 protein expression. Proliferating tumor cells show increased expression of specific lncRNAs, linking noncoding RNA activity to cell cycle state.
Proliferating epithelial populations show coordinated expression of TUG1 alongside proliferation markers, indicating an association between lncRNA activity and cell cycle progression.
Together, these results demonstrate that whole transcriptome FFPE profiling captures both coding and noncoding features of tumor biology across subtype and cell state.
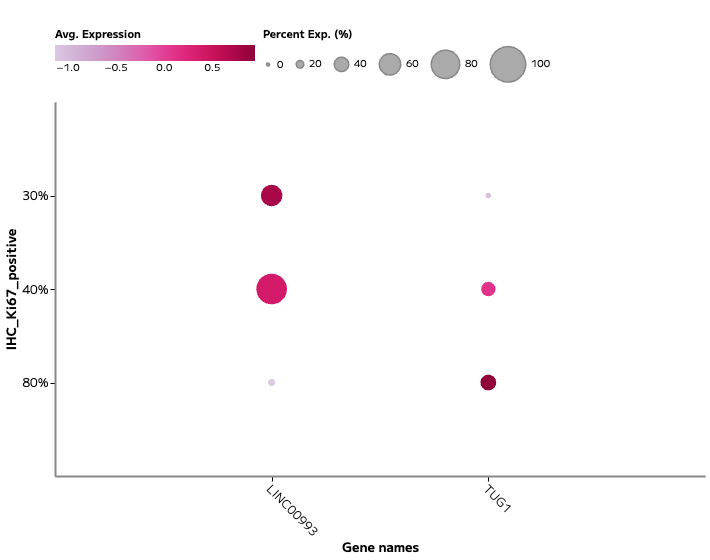
Figure 2: Subtype-associated lncRNA expression across breast tumor cell states.
To further connect lncRNA expression with functional and clinical measures of proliferation, lncRNA expression was assessed using Ki67 positivity, as determined by prior immunohistochemistry protein staining.
Dot plot showing expression of LINC00993 and TUG1 across annotated cell populations. LINC00993 is enriched in luminal epithelial populations, while TUG1 is elevated in TNBC and proliferating epithelial populations. Dot size indicates the percent of cells expressing each transcript and color indicates average expression.

Dr. Ebru Boslem
ANZ Market Manager - Research Genomics