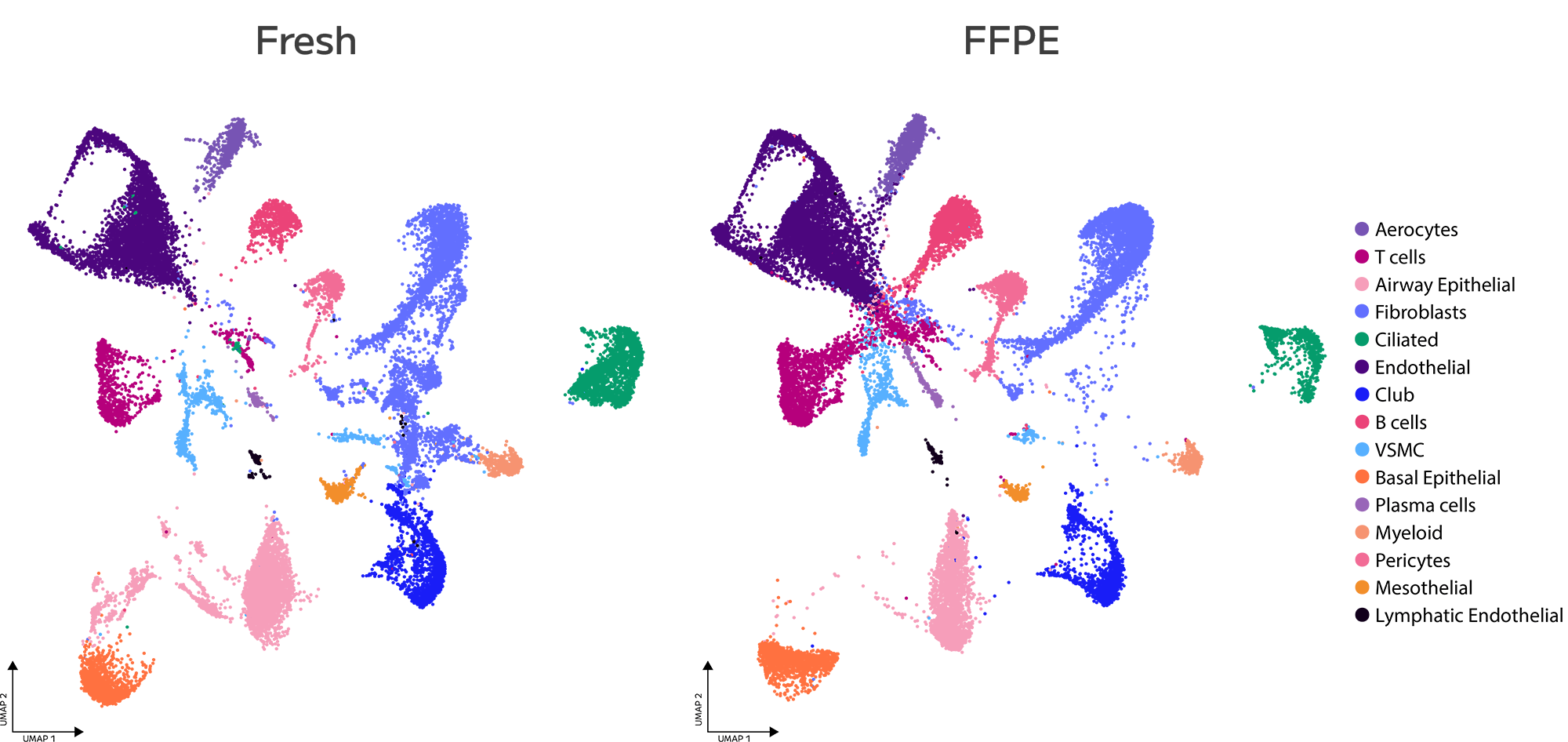TrueAmp Polymerase Mix
/in NGS, Twist/by Harshita Sharma
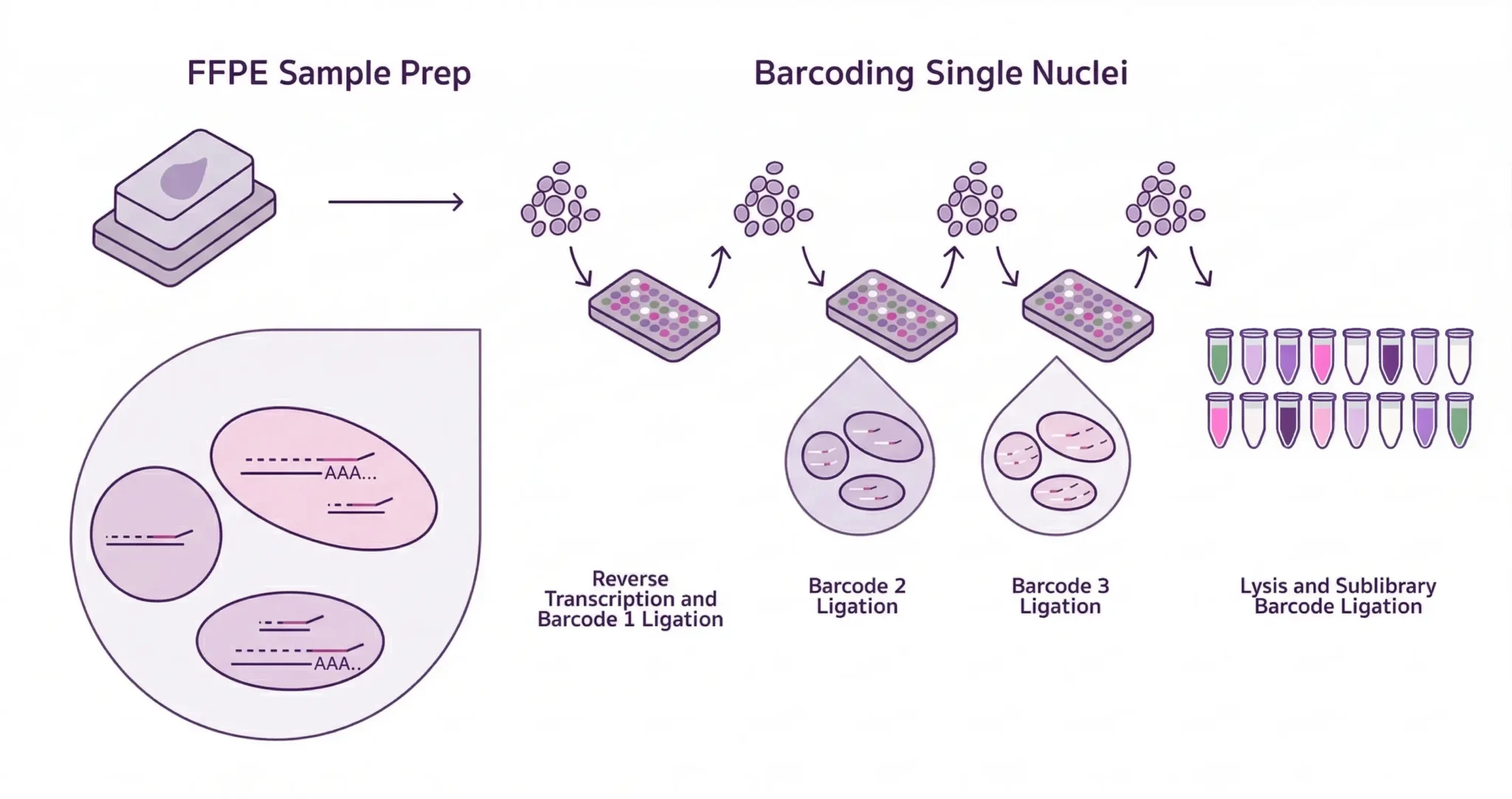
The Twist TrueAmp Polymerase Mix is an engineered amplification reagent purpose-built for NGS library workflows — not adapted from qPCR or cloning applications, but designed from the ground up for the demands of complex library amplification. At its core is a proofreading, high-processivity DNA polymerase formulated to deliver uniform coverage across the full GC spectrum, from AT-rich to GC-rich targets, while minimising the amplification-induced errors and homopolymer slippage that compromise variant calling accuracy. Whether you’re amplifying pre-capture or post-capture libraries, TrueAmp Polymerase Mix keeps bias low and your data trustworthy.
Delivered as a single-tube 2× mastermix with aptamer-enabled hot start, TrueAmp Polymerase Mix is stable at room temperature and ready for automated liquid handling — reducing setup complexity, minimising handling errors, and supporting same-day or overnight hybridisation workflows alongside Twist target enrichment kits. It is validated end-to-end with Twist library preparation and enrichment reagents, making it a natural fit for laboratories already running Twist workflows who want consistent amplification performance without introducing an unvalidated variable. As your authorised Australian distributor, Decode Science can advise on integration into your current library prep and enrichment protocols.
But... Why Choose TrueAmp Polymerase Mix?
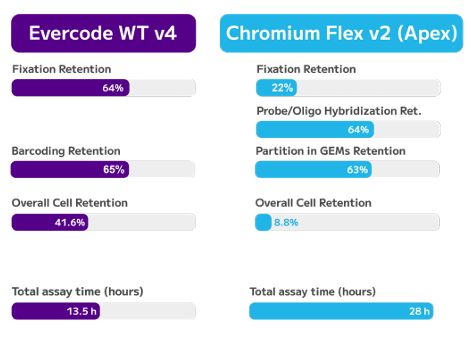
Sustaining coverage uniformly across Full GC Spectrum (5 – 95% )
High Sensitivity & Consistent Yields From Low-Input Libraries
Reduced Homopolymer Slippage and Deamination Errors
Automation-Ready Hot Start in a Simplified Single-Tube Format
Product Data: Performance Across the Metrics That Define Library Amplification Quality
Low-Bias Amplification Across GC Extremes
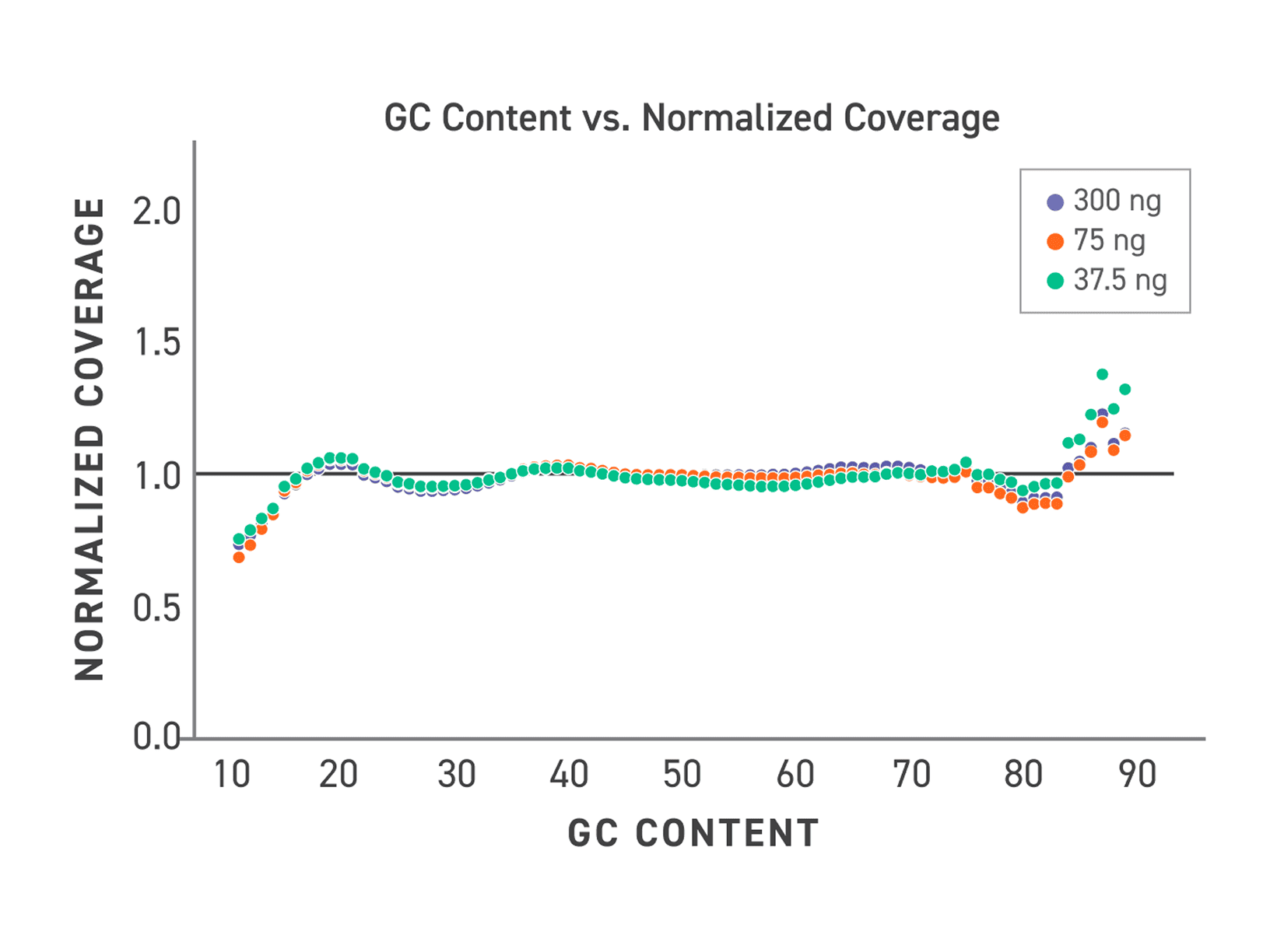
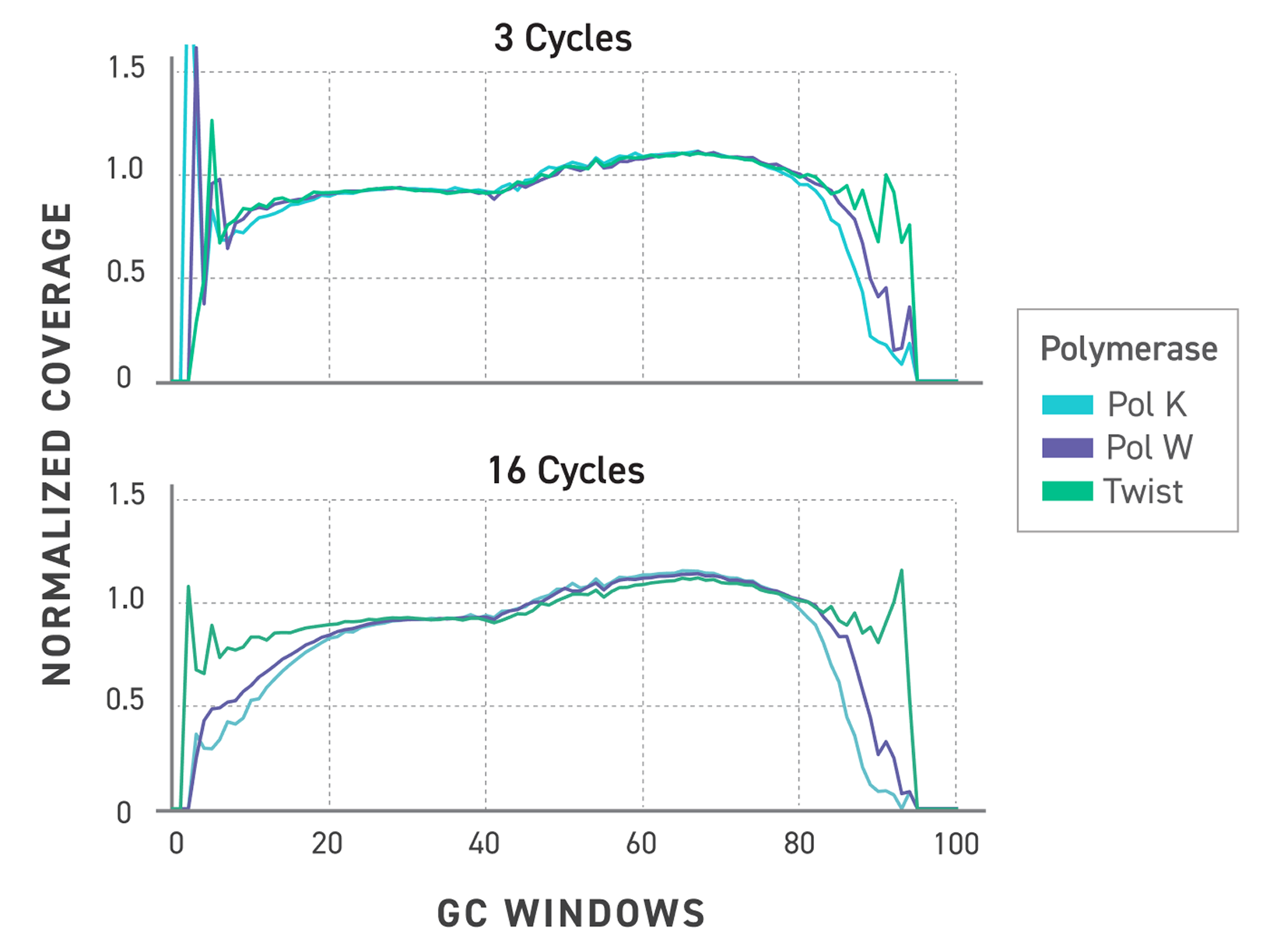
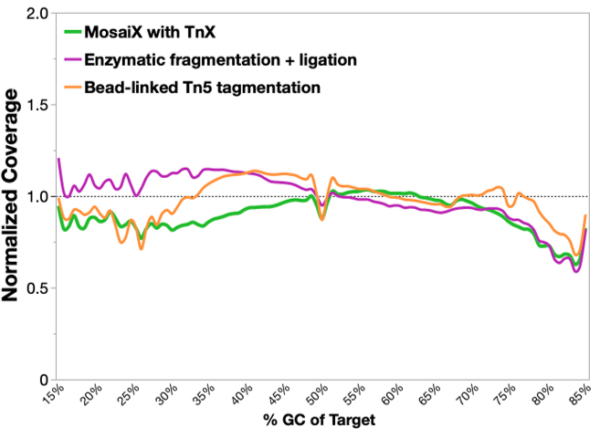
TrueAmp Polymerase Mix sustains normalised coverage across the full GC content range — including AT-rich organisms such as C. difficile and GC-rich organisms such as B. pertussis — where other polymerases show progressive coverage dropout. This performance is maintained as PCR cycle number increases, making it reliable for both low-cycle pre-capture amplification and higher-cycle post-capture workflows.
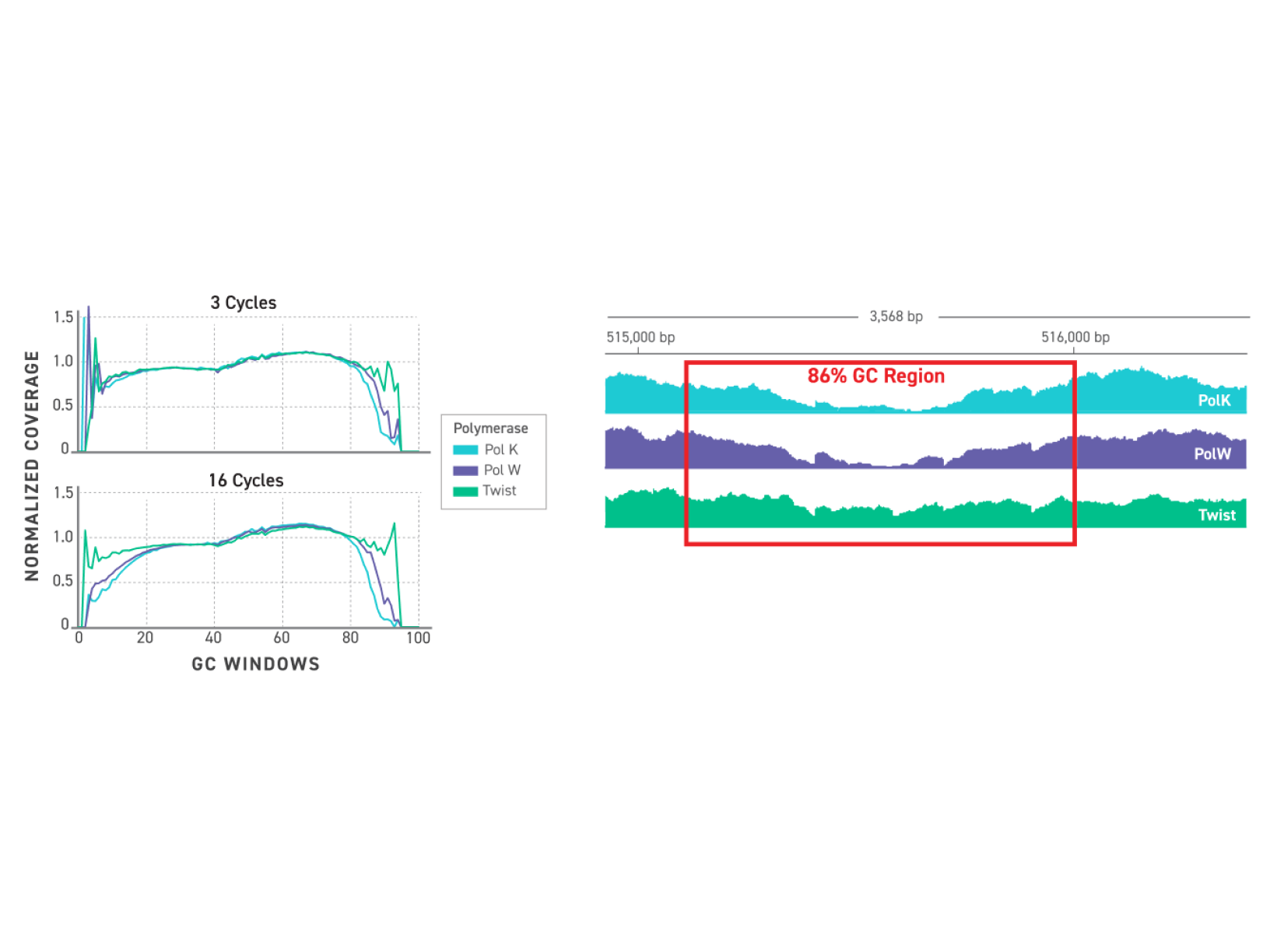
Figure 1. GC-normalized coverage; AT-rich C. difficile and GC-rich B. pertussis.
Efficient Yields at Low Input in Fewer Cycles
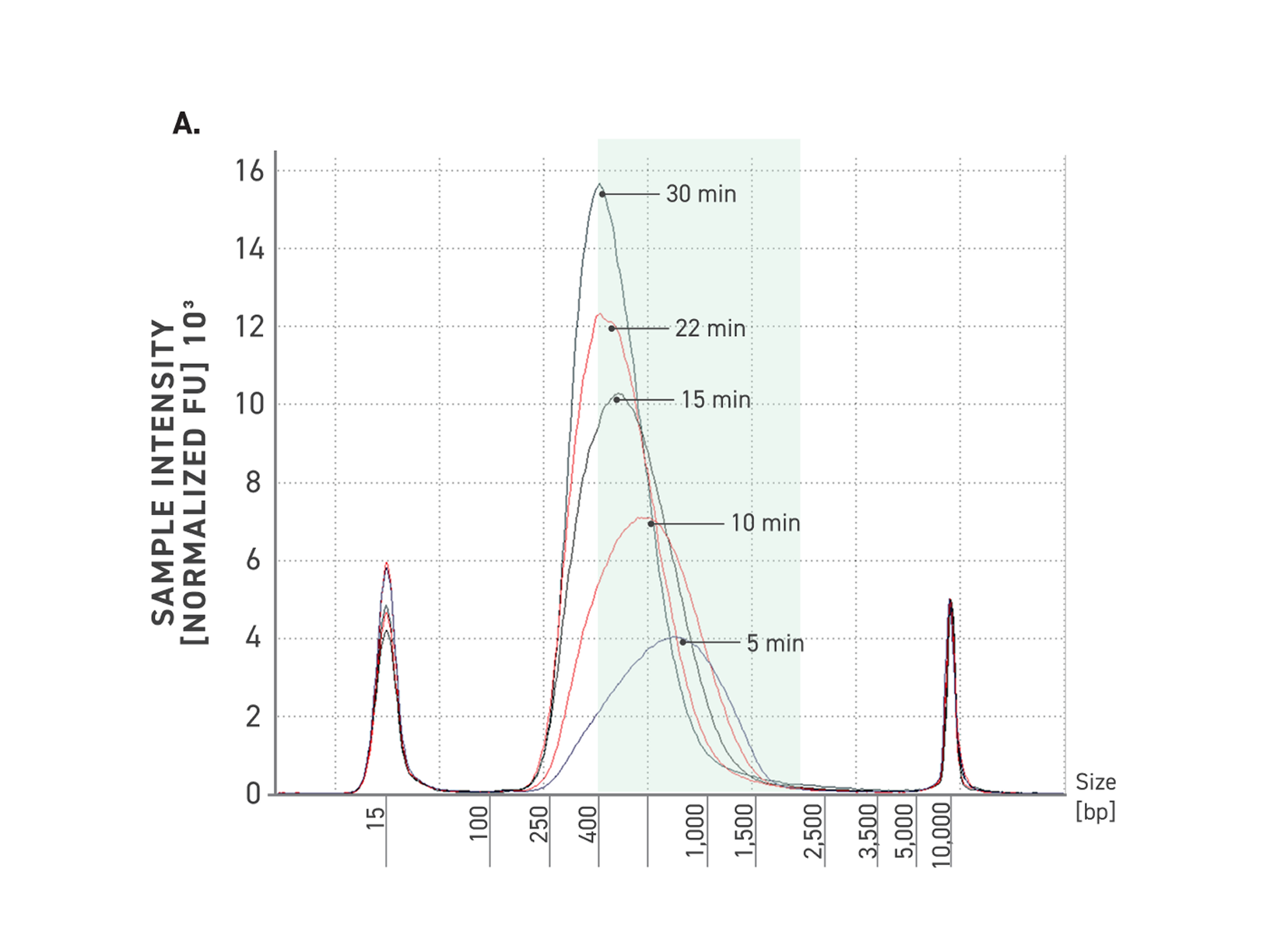
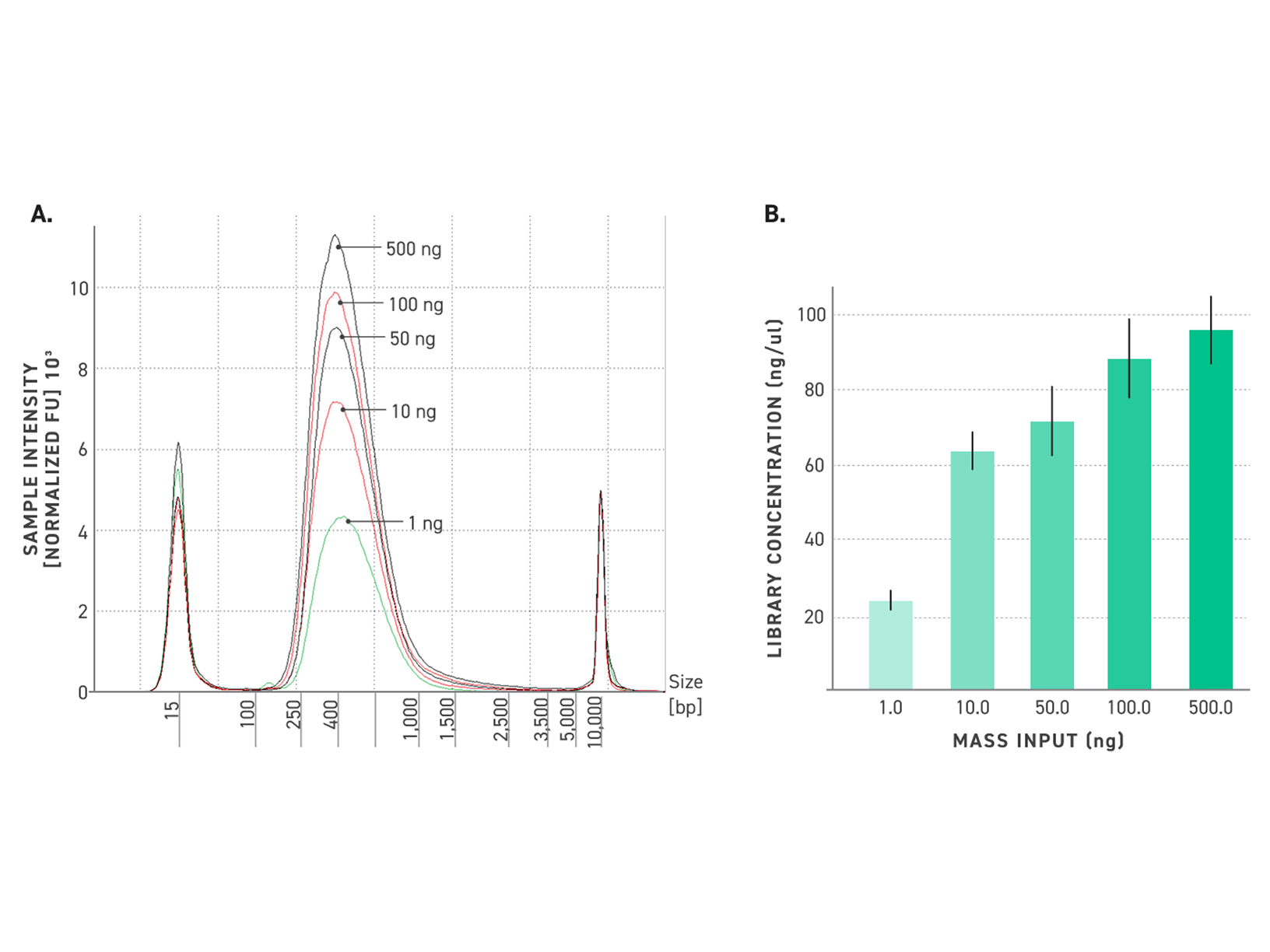
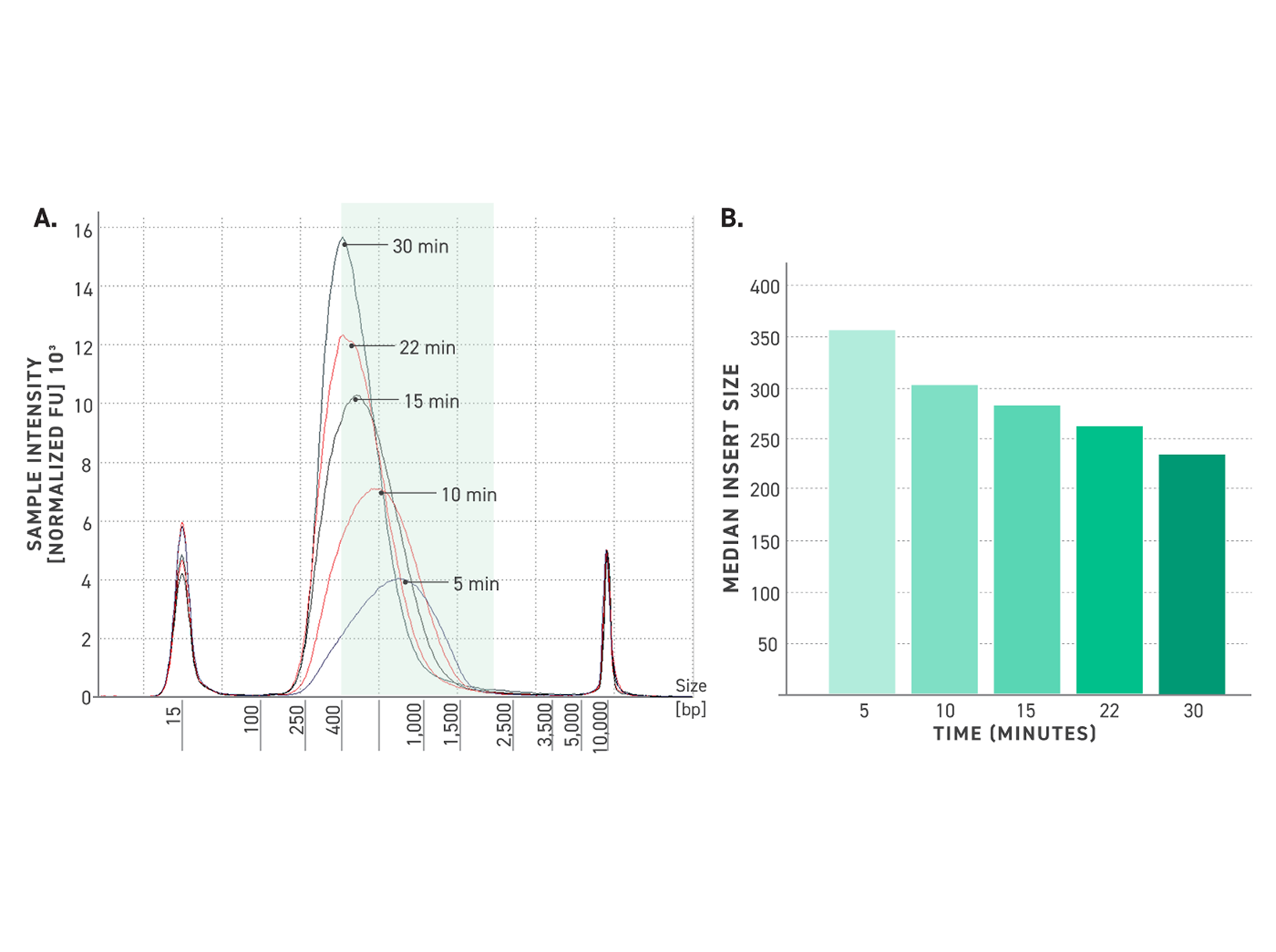
Improved amplification efficiency means TrueAmp reaches yield targets in fewer PCR cycles than standard NGS polymerases. Fewer cycles directly reduces the accumulation of duplicate reads and cycle-induced artefacts — preserving library complexity and sequencing efficiency, particularly from low-yield post-capture libraries where every cycle counts.
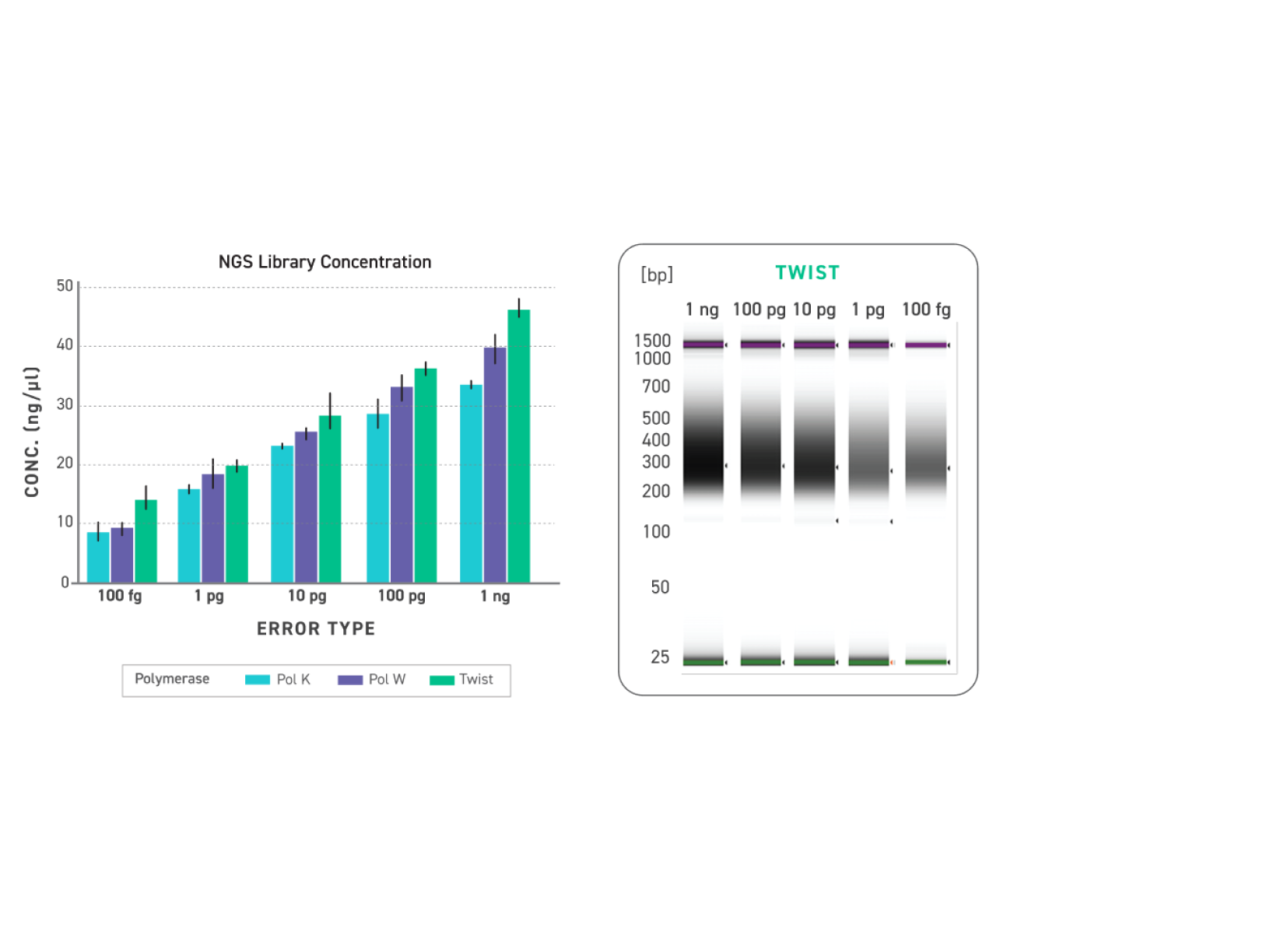
Figure 2. Serial dilutions from 1 ng to 100 fg, Qubit and TapeStation QC.
Reduced C→T Misincorporation and Homopolymer Slippage
Engineered proofreading activity measurably reduces C→T misincorporations introduced by cytosine deamination — one of the most common sources of false positive variant calls in NGS data, and particularly prevalent in FFPE-derived libraries. Improved homopolymer fidelity further reduces a known source of indel artefacts in repeat-containing regions, supporting cleaner variant calls across the full spectrum of library types.
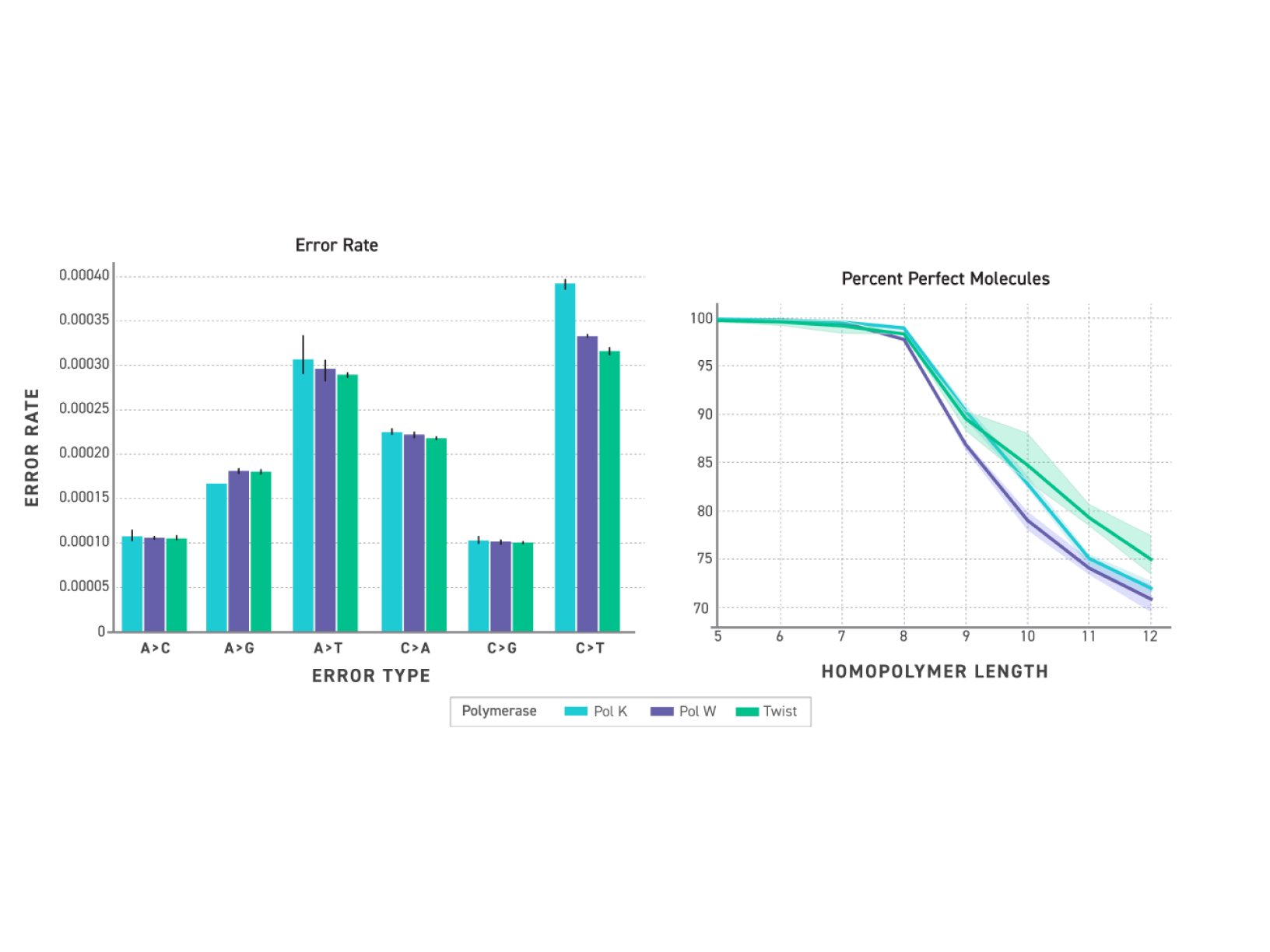
Figure 3. Substitution rates after >7M base incorporations; homopolymer readout using Twist clonal plasmids.
Bead-Tolerant PCR for Reliable Post-Purification Amplification
Magnetic beads used for library purification and size selection can carry over into PCR reactions and inhibit amplification, reducing yields and introducing variability. TrueAmp Polymerase Mix is formulated to maintain robust performance in the presence of magnetic bead carryover — removing a common source of batch-to-batch inconsistency in post-purification amplification steps.
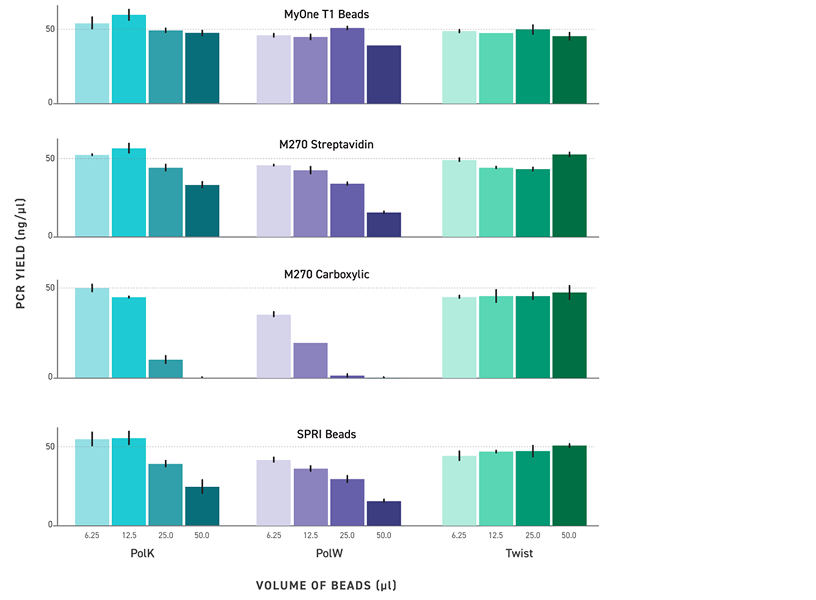
Figure 4. Tolerance to Paramagnetic Beads During PCR. PCR reactions were spiked with 6.25 μl, 12.5 μl, 25 μl, and 50 μl of MyOne T1 Beads, M270 Beads (Invitrogen), and Twist DNA Purification Beads. Reactions were purified post bead-removal and quantified on Qubit dsDNA Broad Range Assay.
Why It Matters to You?
Because Amplification Bias Is One of the Last Variables Most Labs Think to Interrogate
When sequencing results are inconsistent or coverage is uneven, the first instinct is usually to look at sample quality, capture chemistry, or bioinformatics. Amplification polymerase is rarely the first variable examined — but for libraries with challenging GC content or low input, it is often where the problem originates.
TrueAmp Polymerase Mix is most relevant in these contexts:
Target enrichment and exome sequencing —
GC-extreme regions are routinely underrepresented in enriched libraries. TrueAmp’s uniform coverage across 5–95% GC content ensures those targets are captured and sequenced with the same confidence as GC-neutral regions.
Post-capture PCR from low-yield libraries —
After hybridisation capture, the library is often at its most precious and most limited in quantity. An amplification reagent that achieves yield targets in fewer cycles with less bias preserves the complexity and integrity of that material.
Somatic variant detection and low VAF calling —
Reduced C→T misincorporation and homopolymer slippage directly improves the signal-to-noise ratio for low-frequency variant detection — critical for tumour heterogeneity studies, liquid biopsy, and minimal residual disease monitoring.
FFPE-derived libraries —
Deamination damage in FFPE DNA generates artefactual C→T changes that a standard polymerase will faithfully amplify. TrueAmp’s engineered proofreading reduces this signal, preserving the accuracy of variant calls from archival material.

Chris Wicky
Clinical Genomics Manager - ANZ & Country Manager - NZ
Related Products
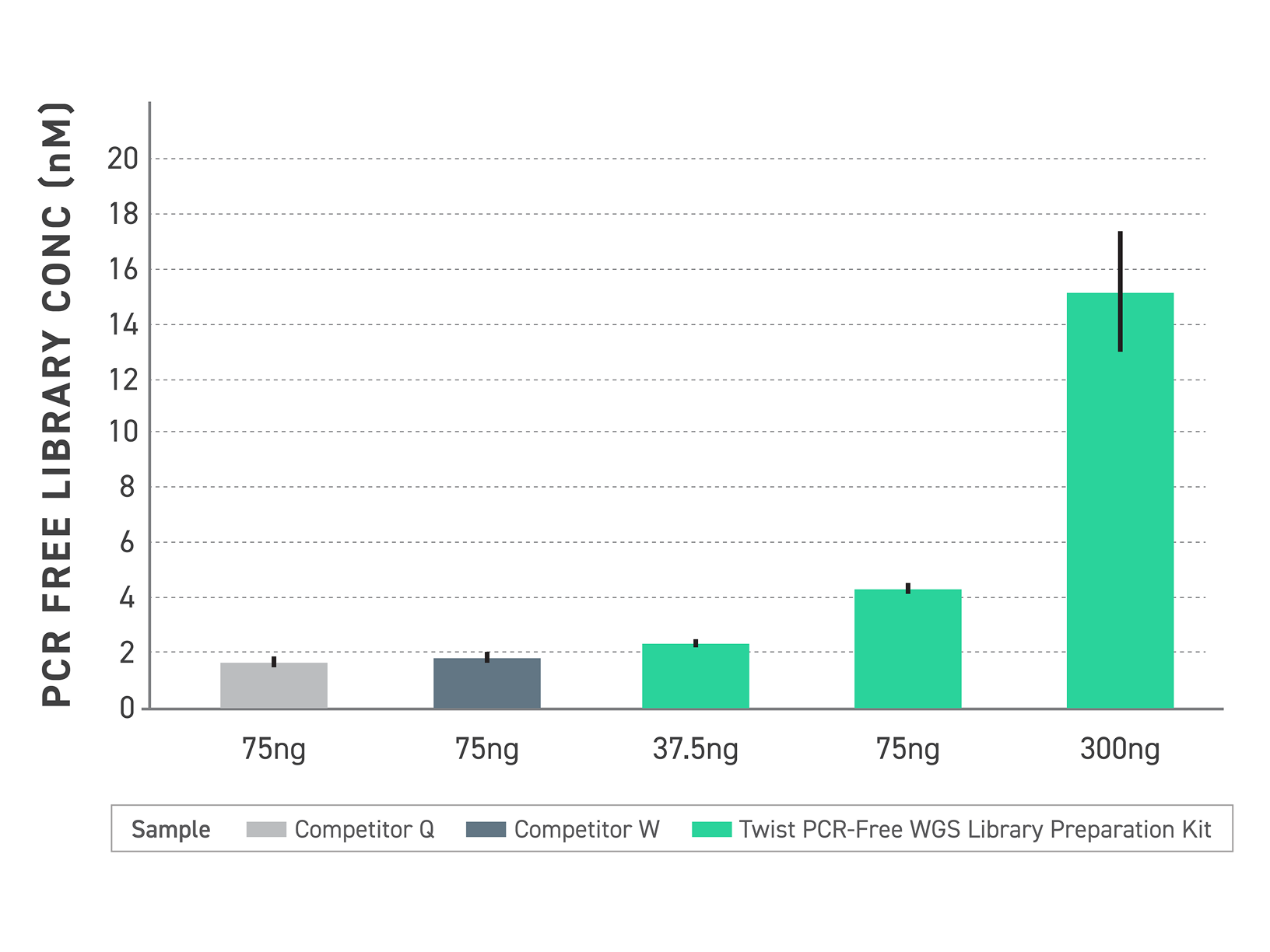
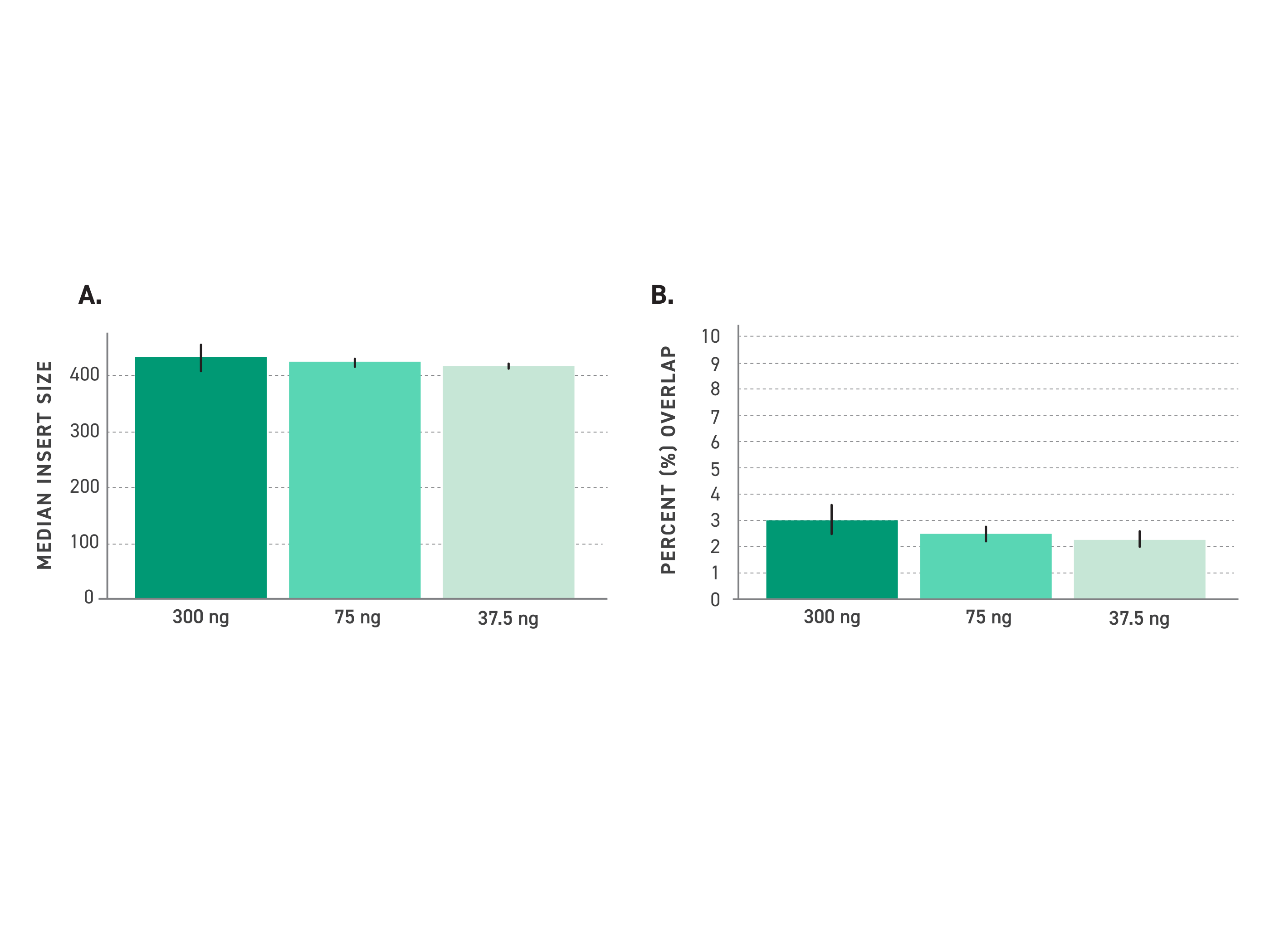
Twist PCR-Free WGS Library Preparation Kit
Bias-free whole genome libraries from high-quality input DNA, no amplification required
Twist Custom NGS Panels
Design and order target enrichment panels tailored to your gene list or genomic region of interest
Twist Exome 2.0
Comprehensive exome capture panel with proven uniformity across canonical and difficult targets
Resources
Unlock with quick sign up!
FAQs
What is TrueAmp Polymerase Mix and how is it different from standard PCR polymerases?
TrueAmp Polymerase Mix is an engineered proofreading, high-processivity DNA polymerase formulated specifically for NGS library amplification. Unlike standard PCR enzymes optimised for single-amplicon applications, TrueAmp is designed to amplify complex NGS libraries with uniform GC coverage, high fidelity, and consistent yields — minimising the bias and artefacts that standard enzymes introduce when applied to library amplification.
What is the minimum input TrueAmp Polymerase Mix can handle?
TrueAmp Polymerase Mix generates consistent, high-yield libraries down to 100 pg input or below. Contact Decode Science for guidance on cycle number optimisation at very low inputs.
Is TrueAmp Polymerase Mix compatible with my existing Twist library prep protocol?
TrueAmp Polymerase Mix is validated end-to-end with Twist library preparation and enrichment reagents. It is compatible with pre-capture and post-capture amplification steps and with same-day and overnight hybridisation workflows alongside Twist target enrichment kits. Contact Decode Science to confirm compatibility with your specific protocol configuration.
Can TrueAmp Polymerase Mix be used with non-Twist library prep kits?
TrueAmp Polymerase Mix has been validated with Twist library prep and enrichment reagents. For use with other library prep systems, contact Decode Science to discuss compatibility and any protocol adjustments that may be required.
Does the hot start format affect performance?
No. The aptamer-enabled hot start provides room temperature stability for setup without compromising PCR performance. It eliminates the need for ice or cold block handling during reaction assembly, simplifying manual workflows and supporting automated liquid handling integration.
Is TrueAmp Polymerase Mix suitable for FFPE-derived libraries?
Yes. TrueAmp’s engineered proofreading activity reduces C→T misincorporations caused by cytosine deamination — a common artefact in FFPE-derived DNA — making it particularly well-suited to post-capture amplification from FFPE libraries where deamination damage is elevated.
What sequencing platforms are TrueAmp Polymerase Mix libraries compatible with?
TrueAmp Polymerase Mix produces libraries compatible with Illumina sequencing platforms. Contact Decode Science for platform-specific guidance.
Talk to Us About TrueAmp Polymerase Mix
We only need these information to serve you better. Decode Science respects your privacy and will never spam you with unrelated content.