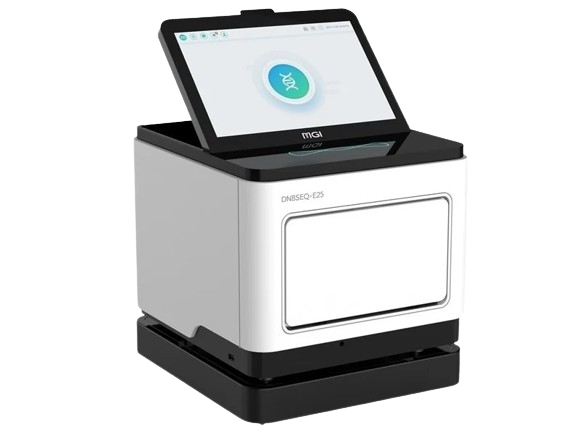
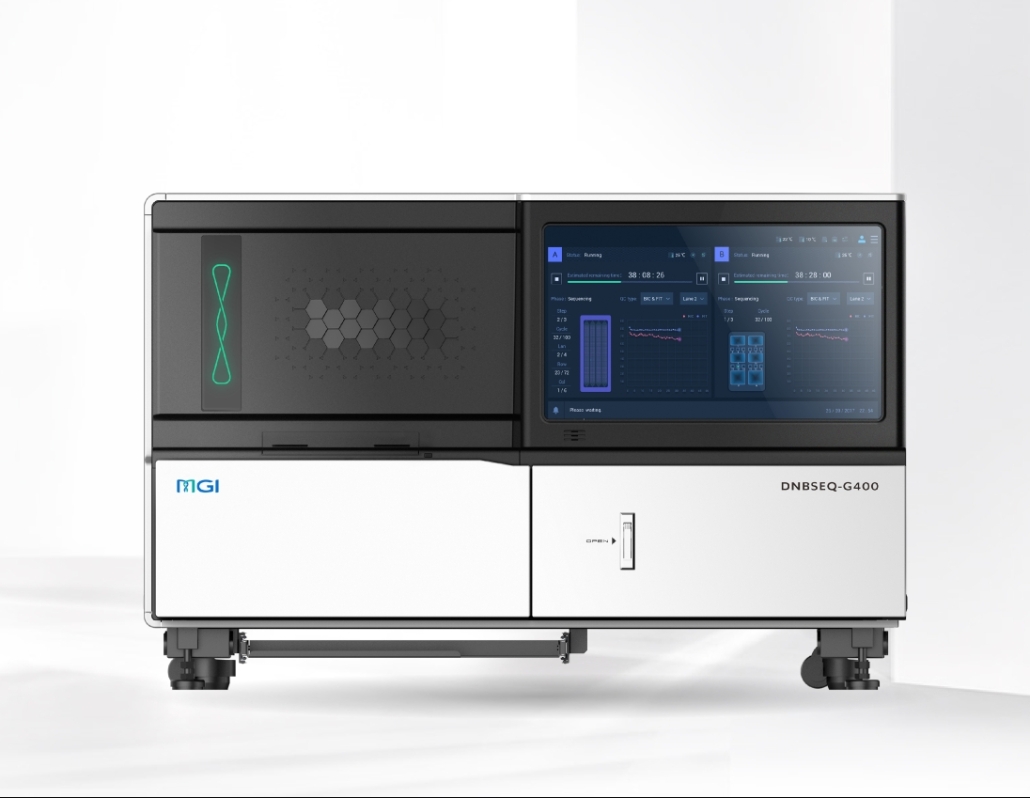
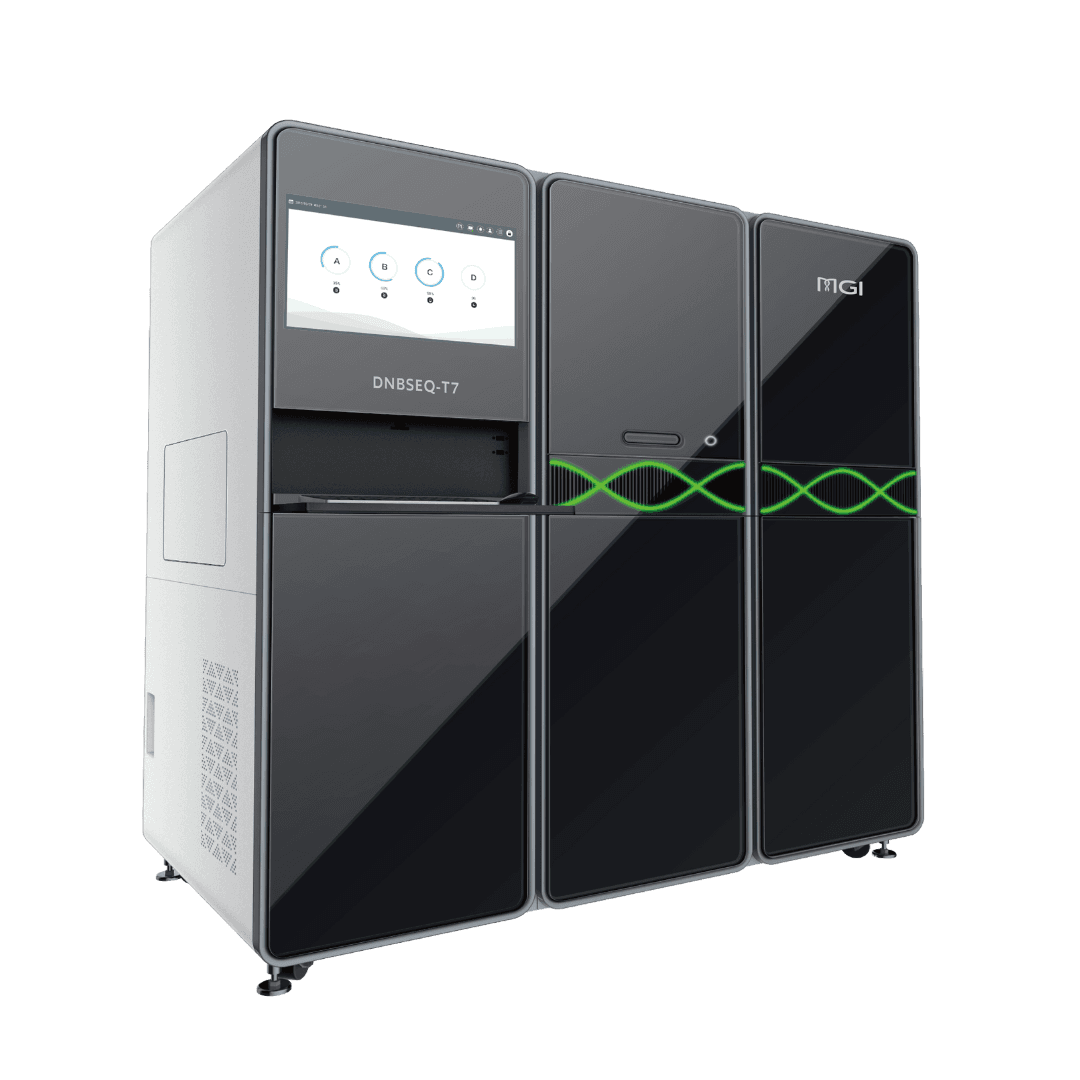
DNBSEQ T7+
/in MGI, Partners, Sequencing Platforms/by Harshita Sharma
T7+ is an integrated ultra-high-throughput sequencing platform built for labs that demand speed, accuracy, and scale. Leveraging MGI’s DNBSEQ™ Technology and SM2.0 biochemistry, it delivers over 14Tb of high-quality sequencing data in 24 hours, making large-scale genomics projects—from population studies to clinical pipelines—both feasible and efficient. Its 7-in-1 modular design automates the entire workflow, reducing hands-on time and minimizing potential errors.
Beyond sheer performance, T7+ is engineered with clinical relevance in mind. Its capacity for up to 35,000 whole-genome sequences annually ensures that your lab can meet high-throughput demands without compromising turnaround times. This efficiency directly translates to faster research insights, improved diagnostic workflows, and smoother integration into daily lab operations, helping you focus on what matters most: accurate, actionable genomic data.
>14 Tb/24h ultra-fast sequencing
7-in-1 automation: prep to analysis
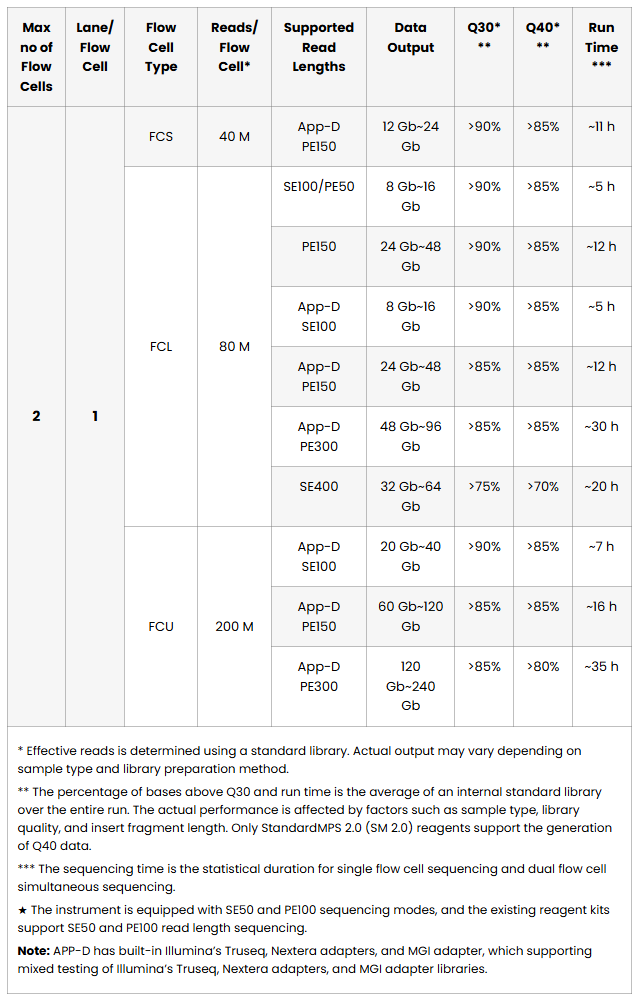
QUAD-Flow Cells: PE150 & PE100 simultaneous
Q40 >90% high-quality output
Ergonomic & modular design
Smart system with diagnostics & self-healing
Omni-smart hub guides workflows
Checkpoint resume for uninterrupted runs
Learn more about how each feature works as you scroll.
Every hour matters when precision, speed, and reliability determine outcomes. T7+ addresses these challenges by combining ultra-high throughput, rapid turnaround, and intelligent automation, so you can focus on results rather than processes.

Ultra-Fast Sequencing:
The proprietary TDI camera and high-density flow cells deliver over 14 Tb of high-quality data in 24 hours, supporting up to 35,000 whole-genome sequences per year. Large-scale projects can now be completed without bottlenecks.

Seamless Automation:
The 7-in-1 workflow integrates DNB preparation, loading, sequencing, waste management, data analysis, and compression. This reduces hands-on time, minimizes errors, and produces ready-to-analyze FASTQ files with Q40 >90%.

Chris Wicky
Clinical Genomics Manager - ANZ & Country Manager - NZ
Unlock with quick sign up!

Smart & Intuitive Operation:
Minimalist design, adjustable screen angles, and the Omni-smart hub guide operations with ease, making daily sequencing as simple as interacting with a smartphone.

Reliable Data Management:
Built-in lossless data compression reduces storage and bandwidth needs by up to 5× without compromising accuracy. Intelligent checkpointing ensures sequencing resumes seamlessly after interruptions.
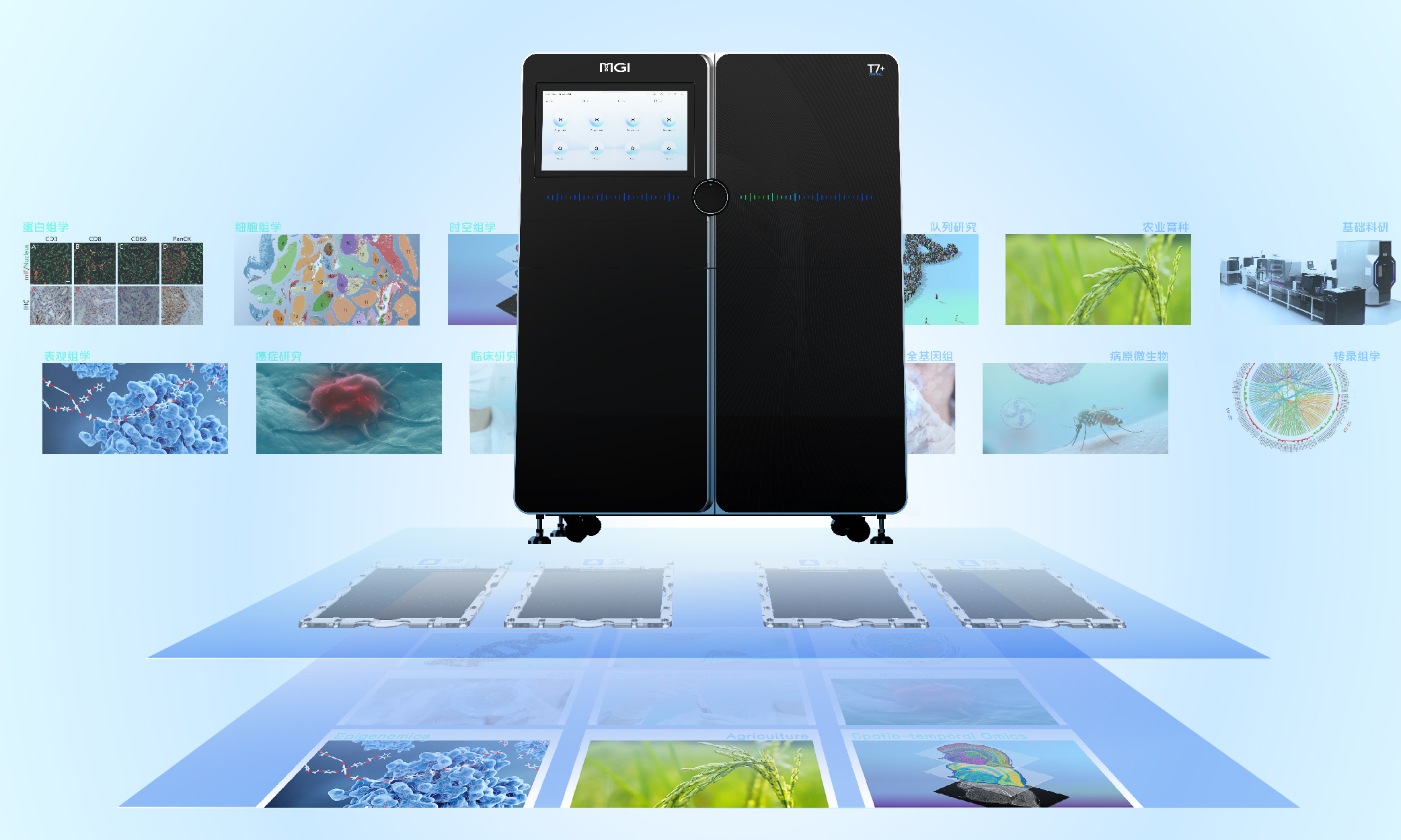

Versatility for Multi-Omics:
T7+ supports WGS, spatio-temporal omics, cell-omics, proteomics, epigenomics, transcriptomics, and more, making it adaptable to evolving research and operational needs.

MGISP-960
MGISP-960 has validated lots of library kits including WES/WGS/RNA and...

DNBelab D series
Highly integrated technology platform which uniquely uses digital microfluidic...

DNBSEQ-T7+RS Reagent Set
A sequencing run starts with the hybridization of a DNA anchor, then ...
A TDI (Time Delay and Integration) camera is a high-performance imaging system used in MGI’s T20×2 ultra-high throughput sequencer to capture fluorescence signals with exceptional sensitivity and speed. Unlike conventional cameras that take static images, a TDI camera continuously scans across the sequencing slide, synchronizing image acquisition with sample movement. This technique integrates multiple exposures of the same area over time, significantly boosting signal strength and reducing noise. In the T20*, the TDI line-scan cameras work alongside a liquid-immersion optical lens and a large field-of-view objective to capture more fluorescence data per unit time with higher resolution. The result is faster, more accurate base identification and greater sequencing throughput — a cornerstone technology enabling MGI’s record-breaking data generation efficiency.
DNBSEQ is MGI’s proprietary sequencing platform based on DNA Nanoball (DNB) technology. Instead of traditional bridge amplification used by other platforms, DNBSEQ amplifies DNA fragments through rolling circle replication, creating dense, uniform DNA nanoballs that are then arrayed on a patterned flow cell. This approach eliminates amplification errors, reduces duplication rates, and enhances signal precision. Combined with MGI’s two-color fluorescence detection and advanced imaging systems, DNBSEQ delivers high-throughput, low-cost, and highly accurate sequencing results. The platform supports a wide range of applications—from whole genome and single-cell sequencing to metagenomics and oncology research—while offering superior data consistency and scalability compared to conventional NGS systems.
Standard MPS 2.0 (SM 2.0) is MGI’s next-generation sequencing chemistry designed to significantly enhance the accuracy and performance of its DNBSEQ™ platforms. By refining enzyme systems, optimizing fluorescent dyes, and improving data interpretation algorithms, SM 2.0 delivers exceptional sequencing quality—achieving over 85% of bases at Q40, or 99.99% base-calling accuracy. These advancements reduce noise, improve signal clarity, and minimize bias from upstream preparation. As a result, researchers gain higher confidence in detecting low-frequency mutations, SNPs, and InDels across applications such as whole genome sequencing, single-cell studies, and microbiome analysis. In essence, SM 2.0 sets a new industry standard for precision, reliability, and data quality in high-throughput sequencing.
T7+ features checkpoint resume technology and proactive fault detection, allowing sequencing to continue without data loss once power or system issues are resolved.
The 7-in-1 integrated workflow minimizes manual intervention—preparation and monitoring are streamlined, so staff can focus on data analysis rather than instrument operation.
Yes. The modular QUAD-Flow Cell system allows independent runs, supporting both small batches and large-scale sequencing without compromising speed or accuracy.
The platform is designed to fit seamlessly into existing operations, with smart guidance, automated data processing, and flexible output formats that simplify downstream analysis.