Smash the limits of single cell sequencing with Parse
/in Seminar/by Harshita SharmaSmash the limits of single cell genomics.
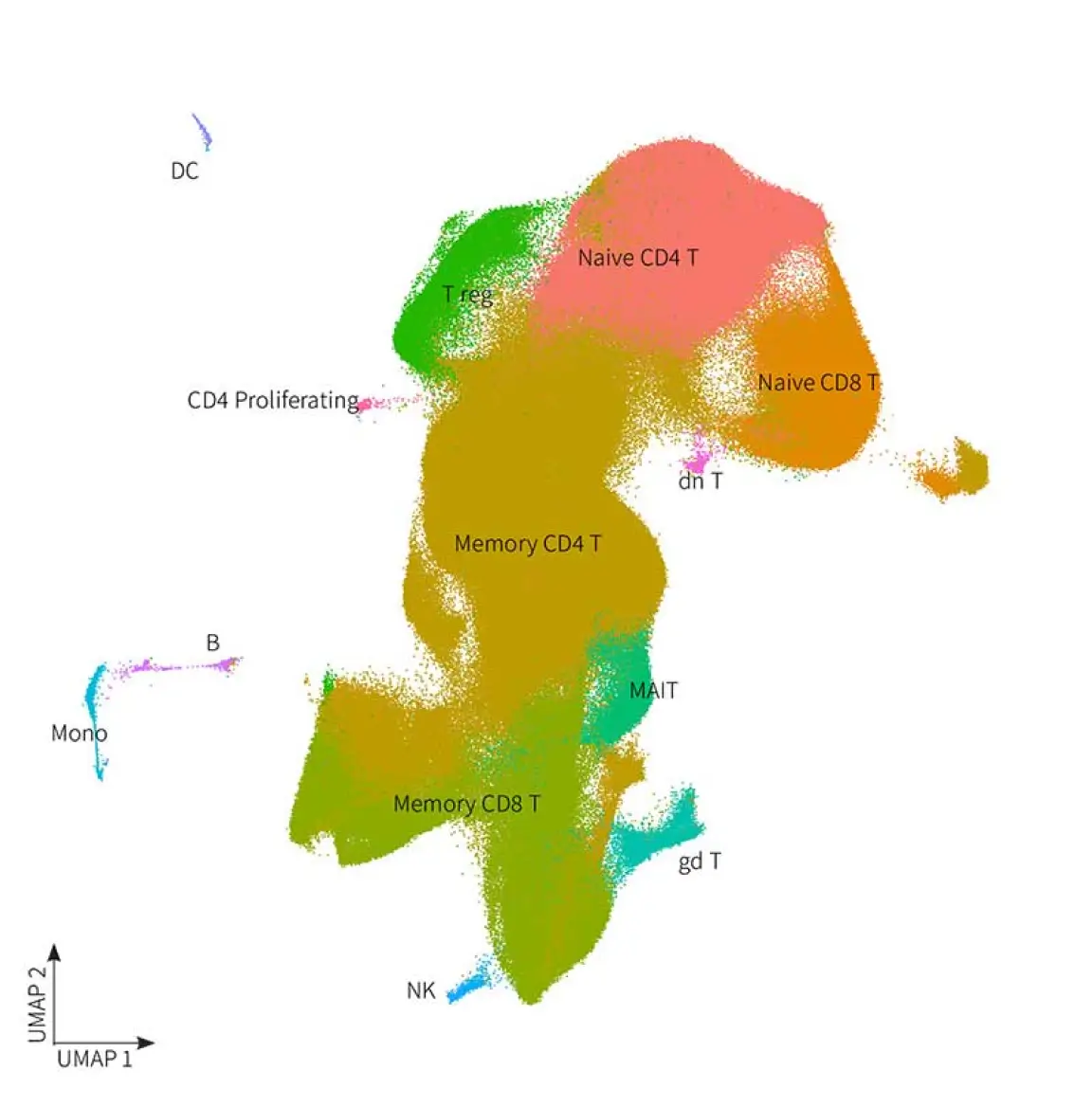
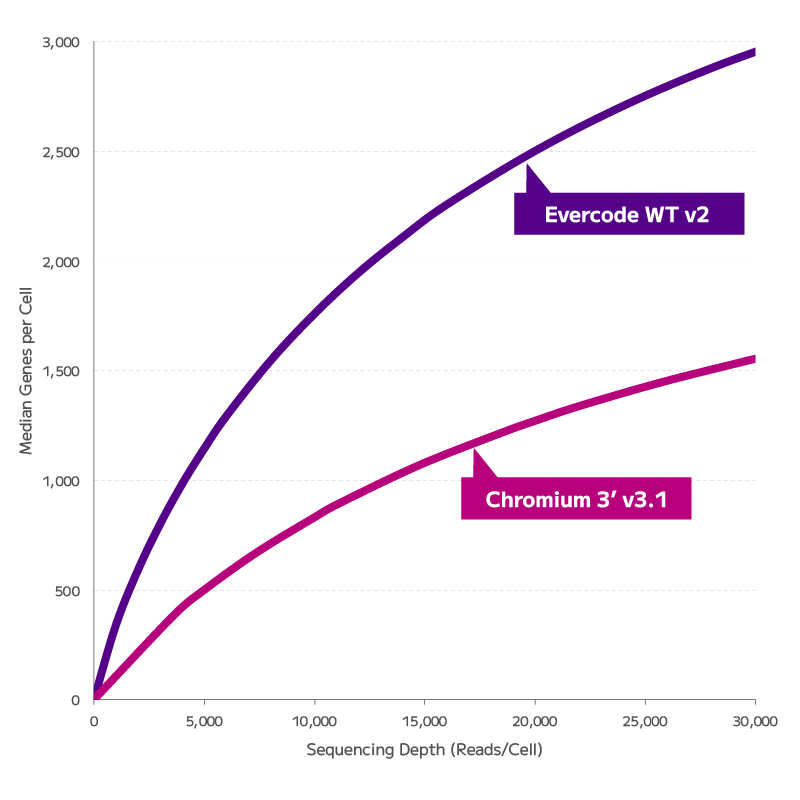
Join us to learn about Parse’s single cell whole transcriptome technology and recently
launched Evercode V3 Kits. More cells, more samples, more clarity.
Combinatorial barcoding technology strips away the limitations and frustrations of yesterday’s single cell approach. It ditches the specialized instrument, freeing you to pursue unprecedented discoveries. Unleash the potential of single cell. Decode Science and Parse Biosciences invite you to a seminar discussing the advances in fixation-based single cell transcriptomics including our V3 chemistry, TCR/BCR kits, CRISPR Detect and Gene Capture.
MEET THE SPEAKER
John received his PhD at Duke University, where he studied cis-regulatory element activation during limb regeneration. He then spent a few years as a postdoc at the University of California, Merced where he studied early organ formation using single-cell genetic and epigenetic approaches. As a Field Application Scientist at Parse Biosciences, John assists scientists with their single-cell genomics experiments, from experimental design, sample preparation, single-cell library workflows, data analysis, and more.
John’s favorite model organism are zebrafish, both embryos and adults. In his free time John enjoys eating spicy food, and dabbles in growing his own hot chili pepper plants.
Brisbane
Date: 8th September 2025
Seminar : 11:30 AM – 12:30 PM – Translational Research Institute Room 3000
Sydney - DAY 1
SEMINAR 1
Date: 9th September 2025
Time: 10 AM – 11 AM
Location:Westmead Institute of Medical Research (hosted by Genomics Core) L2 WIMR: Meeting room C.2.31
SEMINAR 2
Date: 9th September 2025
Time: 2 PM – 3 PM
Location: UNSW (hosted by Ramaciotti) – AGSM Theatre
Sydney - DAY 2
SEMINAR 1
Date: 10th September 2025
Time: 9:30 AM – 10:30 AM
Location: VCCRI (Victor Chang Cardiac Research Institute) (hosted by Innovation Centre), Level 4 Boardroom
SEMINAR 2
Date: 10th September 2025
Time: 12:00 PM – 1 PM
Location: Garvan Institute (hosted by Single Cell Platform) – John Shine Room, Garvan Institute of Medical Research
Canberra
Date: 11th September 2025
Time: 1:00 – 2:00 PM
Location: JCSMR, ANU Seminar Rooms 1+2
Melbourne
SEMINAR 1
Date: 12th September 2025
Time: 11:00 AM – 12:30 PM
Location: WEHI – Genomics Seminar series, Level 7 seminar room
Adelaide
SEMINAR 1
Date: 16th September 2025
Time: 10:00 AM – 11:00 AM
Location: Flinders University Health and Medical Research Building (HMRB)
SEMINAR 2
Date: 16th September 2025
Time: 12:30 PM – 02:00 PM
Location: SAHMRI (South Australian Health and Medical Research Institute)