Parse Single Cell Grant Application – Information Session
/in Seminar/by Harshita SharmaSingle Cell Grant Application - Information Session
Join this webinar to learn about the Parse Biosciences Single-Cell Grant and how researchers can access Evercode™ single-cell technology.
This session is designed for researchers who are new to Parse Biosciences and are interested in applying for the grant.
What we’ll cover:
-
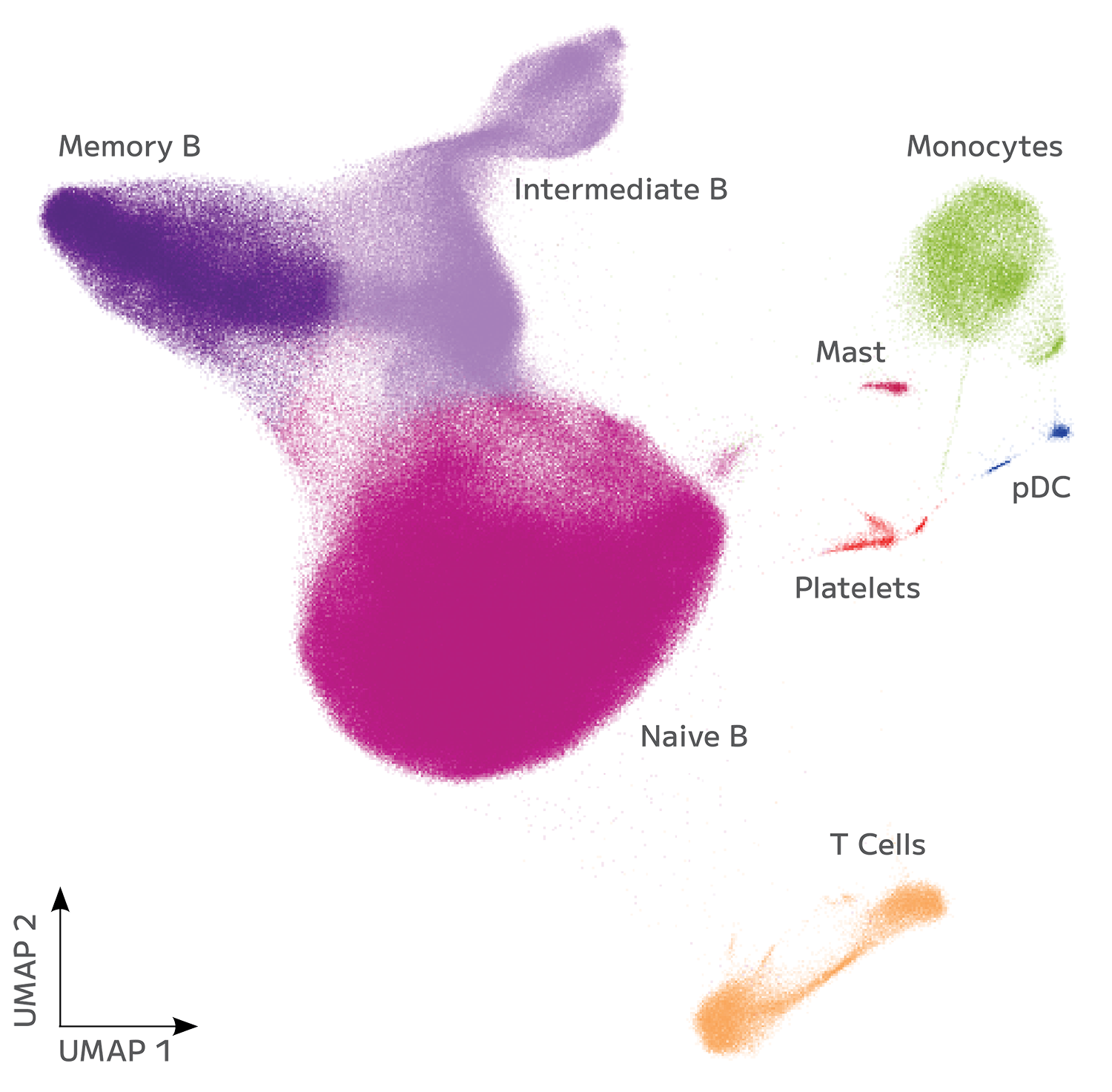
Overview of modern single-cell sequencing approaches
-
Details of the Parse Biosciences grant program
-
Eligibility and evaluation criteria
-
What makes a strong grant application
Attendance is strongly recommended for anyone planning to apply, as the session will include important guidance and updates related to the application process. A recording will be available for registered participants.
Registration is free.