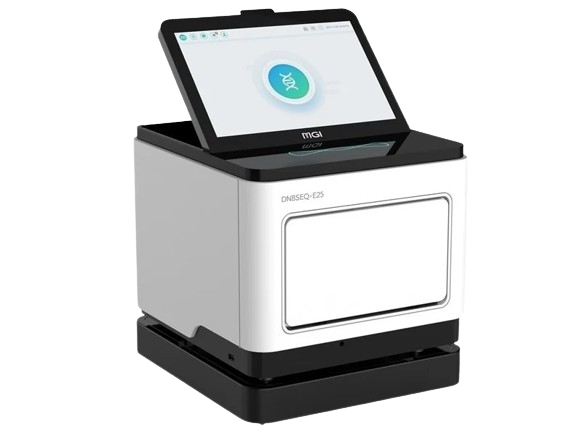
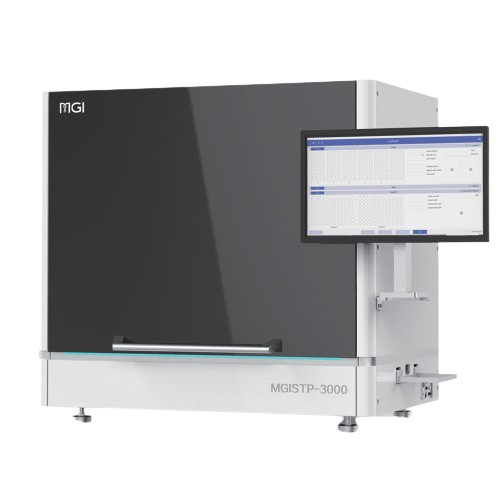
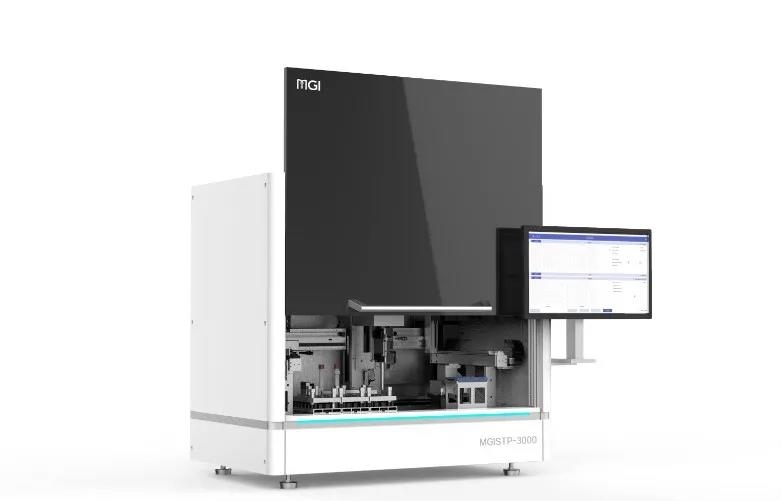
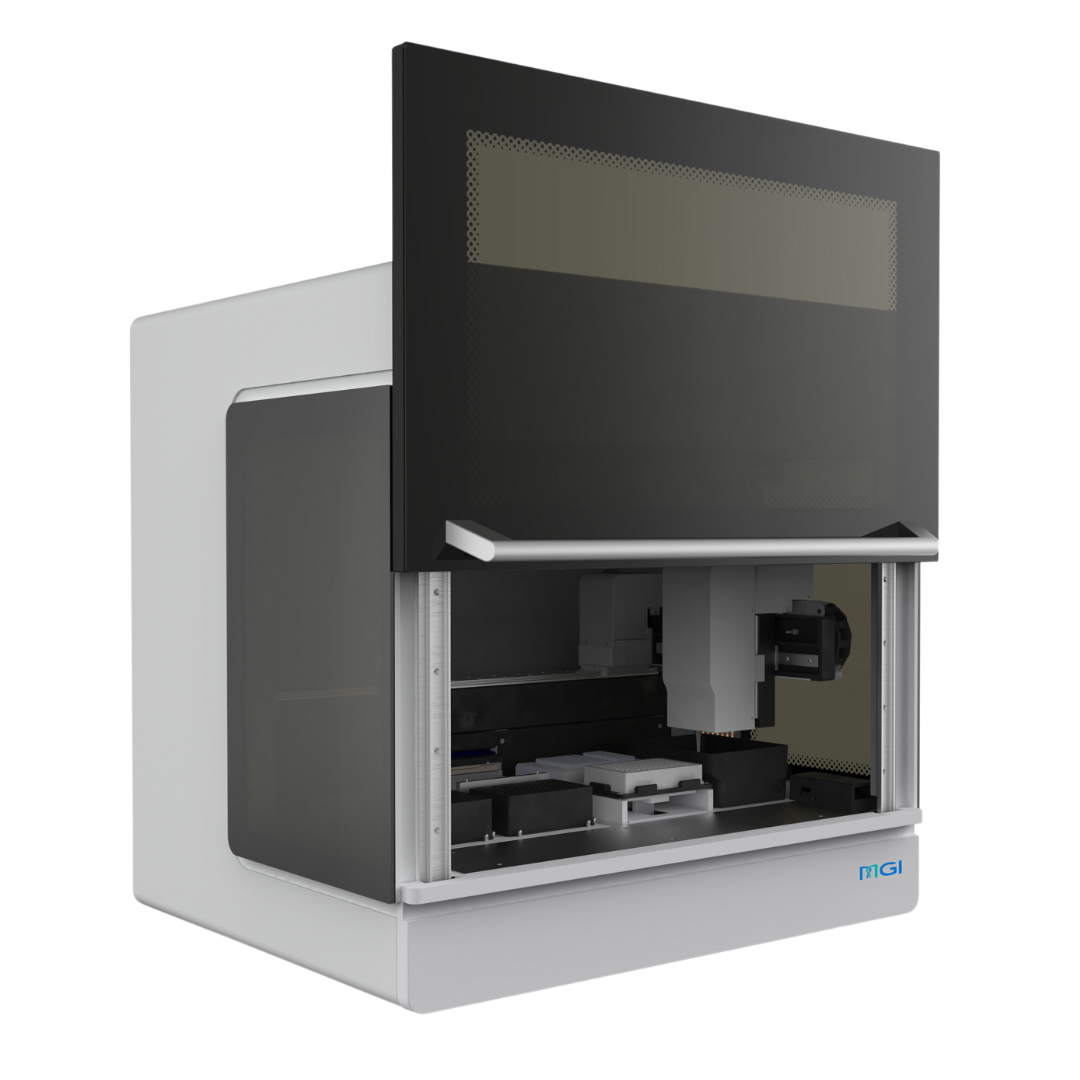
MGISTP-B1000
/in Automated Systems, MGI, Partners, Sample Transfer/by Harshita Sharma
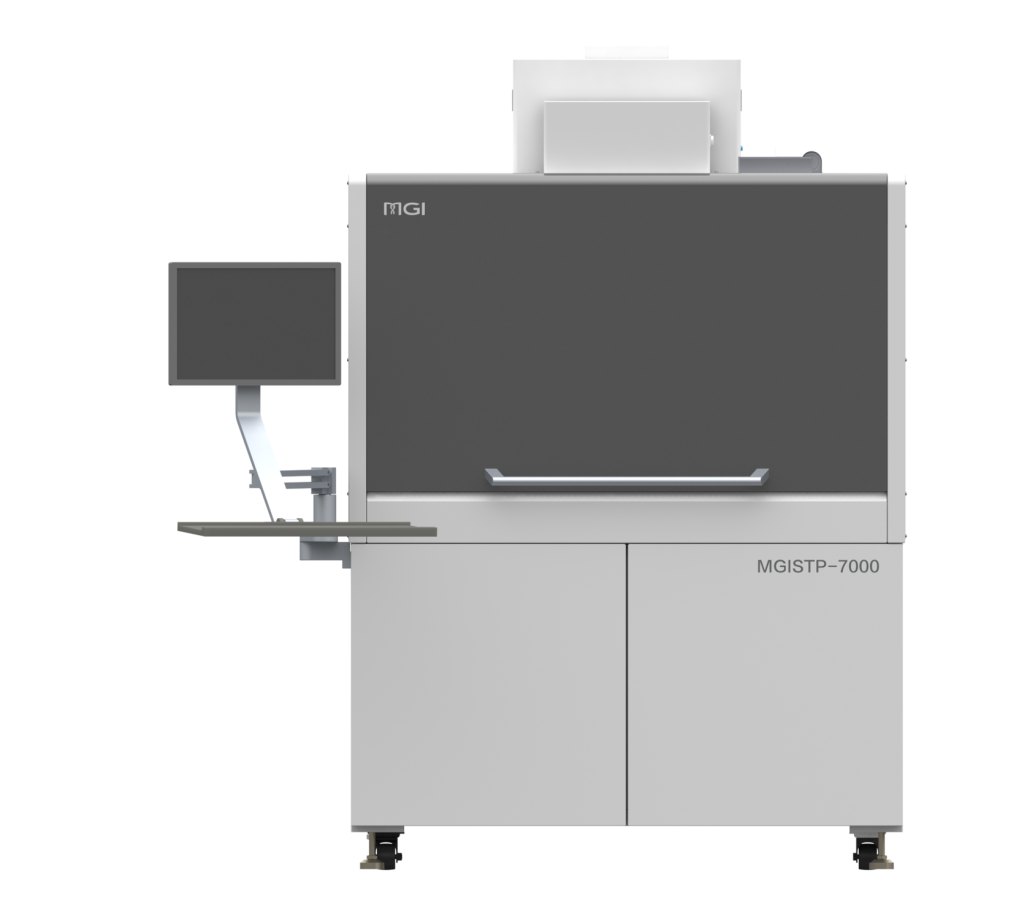
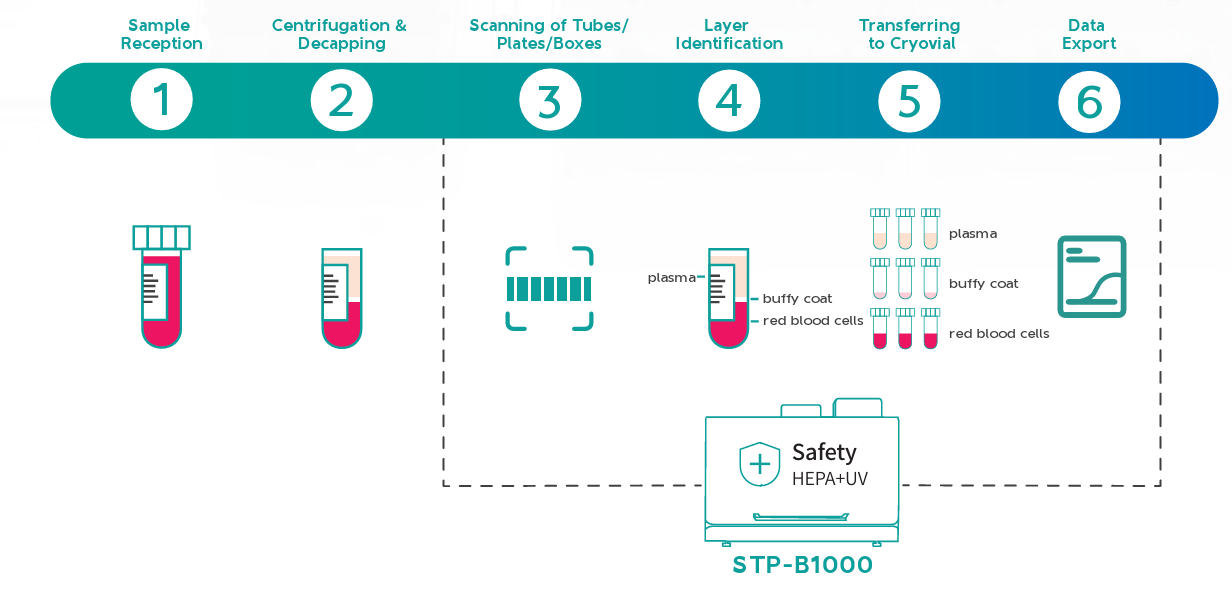
The STP-B1000 is designed for laboratories that require accurate, repeatable separation and transfer of blood components without compromising traceability or efficiency. It precisely recognizes plasma, buffy coat, and red blood cells within centrifuged blood collection tubes and transfers each component with high positional accuracy, reducing manual handling and the risk of cross-contamination. Integrated barcode tracking ensures every transfer remains fully traceable, preventing sample mismatches and data integrity issues in high-throughput workflows.
Operation is intentionally streamlined… users define only the component type, transfer volume, and number of transfers before initiating the process with a single click. This simplicity minimizes training requirements while delivering consistent, standardized blood processing suitable for clinical, biobanking, and research applications.
Precise Layer Positioning
Dual light photography, high-definition camera, built-in self developed image processing
Efficient Buffy Coat Recovery
Spiral aspirate, 3-axis linkage control & recovery rate of 95% or more
Dual Detection Technology
Pressure based liquid level detection (pLLD) & capacitive liquid level detection (cLLD)

Chris Wicky
Clinical Genomics Manager - ANZ & Country Manager - NZ
As the official distributor of MGI in Australia and New Zealand, Decode Science is providing local access to STP-B1000 solutions with region-based technical and application support. Simply talk to me and we can discuss your research needs.
Integrated Scanning, Identification, Transferring
Download Brochure Instantly!

| Indicators | Parameter |
|---|---|
| Pipettor | Pipette Range 1 μL–1000 μL |
| Pipette Accuracy 1 μL: CV≤8%, accuracy: ±10% 50 μL: CV≤1%, accuracy: ±2% 200 μL: CV≤1%, accuracy: ±2% 1000 μL: CV≤1%, accuracy: ±1% | |
| Independent 8-channel Pipettor | |
| Detection Mode | pLLD, cLLD |
| Throughput | 1–192 samples/run |
| Size | 1420 mm (L) × 1010 mm (W) (without door handle) × 1120 mm (H) |
| Weight | ~250 kg |
Contact Decode Science Today
We only need these information to serve you better. Decode Science respects your privacy and will never spam you with unrelated content.