Sample Preparation Kits
/in Bionano, Partners/by Harshita SharmaPRODUCTS
Sample Preparation Kits

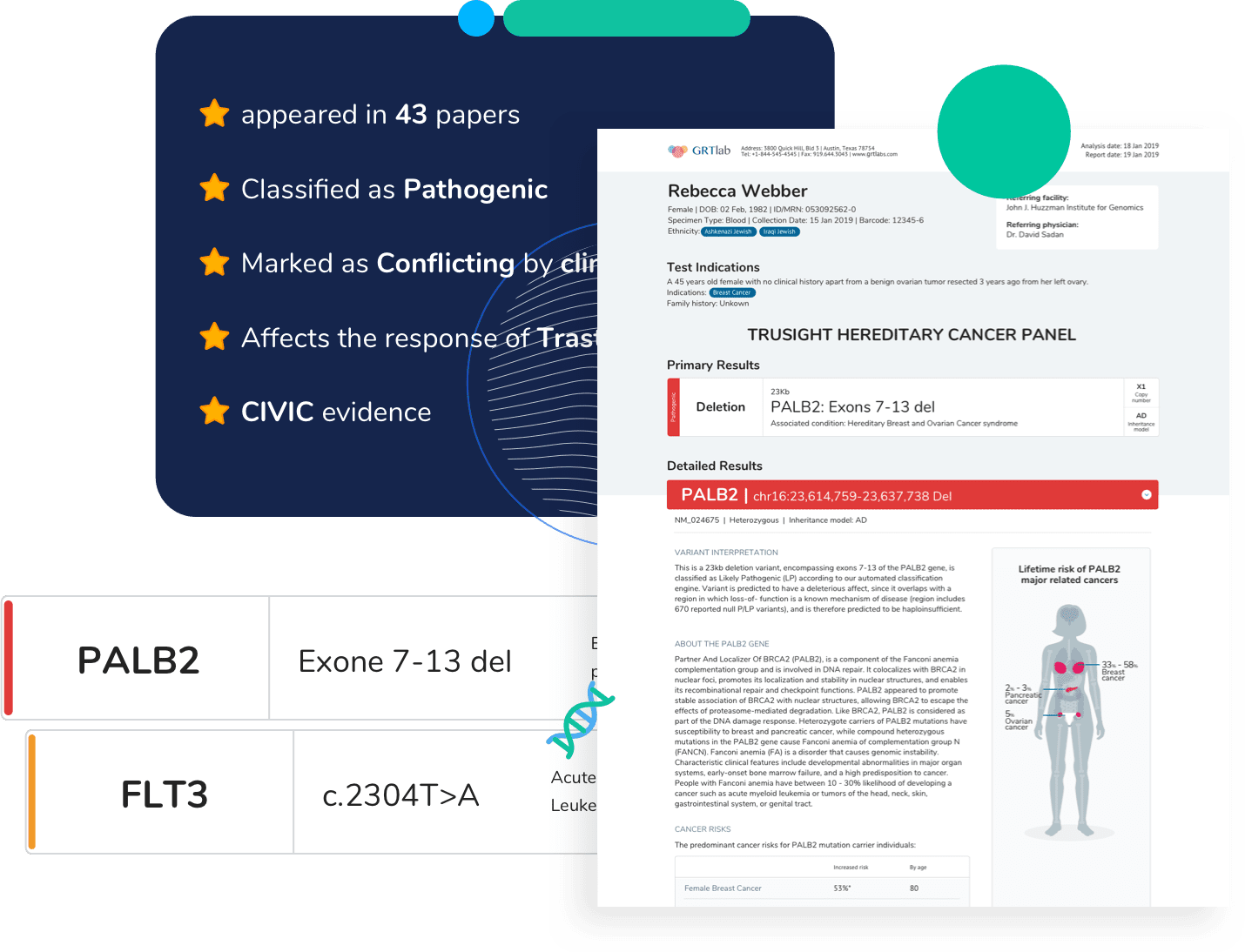
Effortlessly prepare samples for Optical Genome Mapping (OGM) with Bionano’s comprehensive Sample Preparation Solutions, ensuring accurate identification of structural variants.
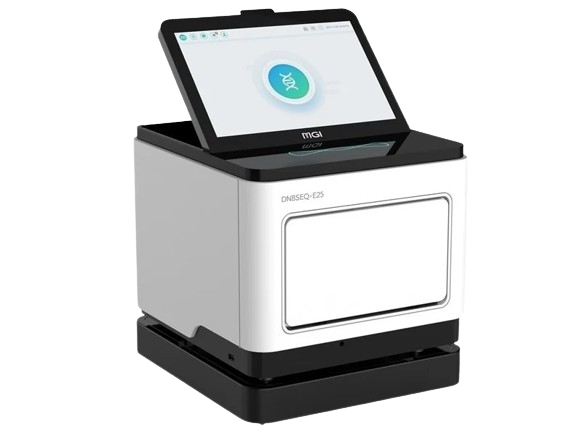
OGM necessitates labeled ultra-high molecular weight (UHMW) DNA, and Bionano offers a diverse range of DNA sample prep kits designed for simple and reliable isolation of UHMW DNA from various sample types. Once DNA isolation is complete, our Direct Label and Stain (DLS) kits can efficiently label DNA for compatibility with the Saphyr system.
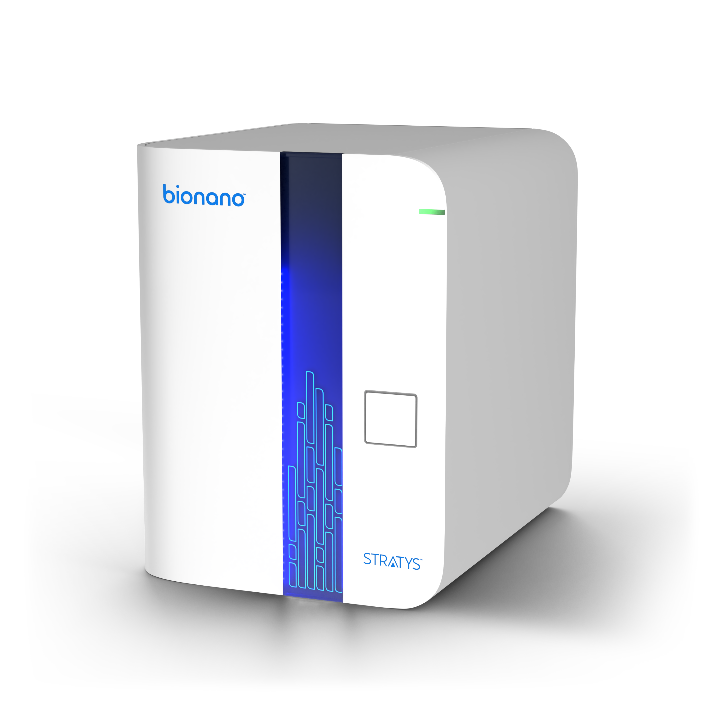
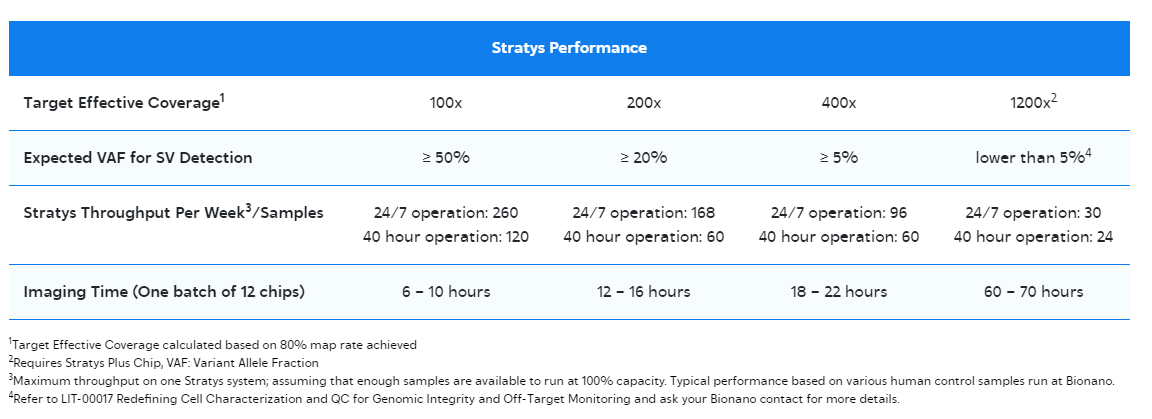
With our Generation 2 Kits, featuring robust enhancements and optimizations to the sample preparation OGM workflow, you can achieve sample-to-answer results in just 3 to 4 days! Experience accelerated and streamlined sample preparation, paving the way for enhanced genomic analysis and structural variant detection.
Attain robust isolation and labeling of Ultra-High Molecular Weight (UHMW) DNA, crucial for visualizing large genomic structural variants.
Bionano’s innovative Solution Phase (SP) technology facilitates the isolation of DNA fragments exceeding 1 Megabase pair (Mbp) in length, consistently yielding average fragments surpassing 230 kilobase pairs (kbp).
To contextualize, while short-read sequencing generates reads up to 150 base pairs (bp) in length, even leading high-accuracy long-read sequencing methods offer read lengths limited to approximately 25 kilobases (kb). Bionano’s SP technology surpasses these limitations, enabling the analysis of genomic structural variants with unprecedented detail and accuracy.

Extract DNA from a diverse array of crucial sample types effortlessly with the latest Bionano SP DNA prep kits.
These kits excel at purifying Ultra-High Molecular Weight (UHMW) DNA from tissues, tumors, bone marrow aspirate (BMA), blood, cells, as well as various plant and animal tissues. This versatility makes Optical Genome Mapping (OGM) applicable across a wide spectrum of studies and applications, including oncology, constitutional genetic disease research, bioprocessing, and general scientific inquiry.
Bionano’s SP kits necessitate 1.5 * 10^6 cells (for blood, cell lines, and BMA) or 10 – 30 mg of tissue as input. Employing a lyse, bind, and wash process along with novel paramagnetic disks, these kits enable the isolation of UHMW DNA in approximately four hours. This streamlined workflow ensures efficient extraction of high-quality DNA, empowering researchers across various fields to unlock the mysteries of the genome with ease.
Boost your workflow throughput by integrating the efficiency of automation.
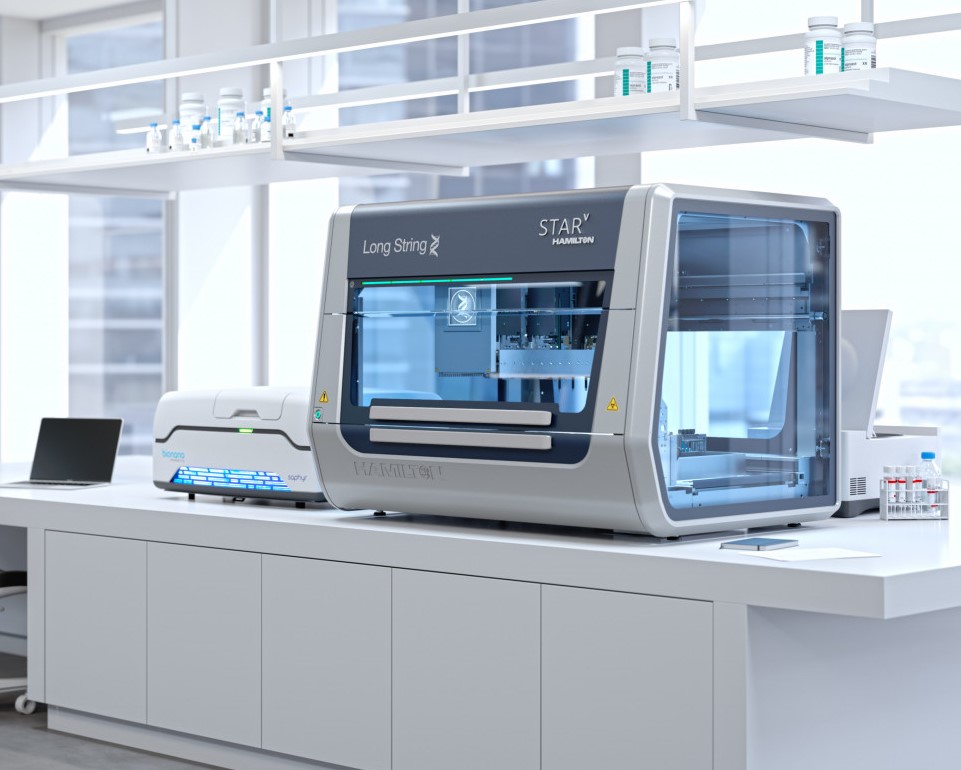
Through a pioneering collaboration with the Hamilton Company, Bionano presents the groundbreaking Long String Star V – the world’s inaugural automated platform tailored for the isolation of Ultra-High Molecular Weight (UHMW) DNA.
The Long String Star V stands as an assay-ready workstation, harmoniously designed to complement Bionano’s G2.LS DNA Isolation kits. With this innovative platform, a single operator can seamlessly extract UHMW DNA from up to 24 samples per day, revolutionizing sample processing efficiency while maintaining high-quality results. Experience unparalleled productivity and precision as you propel your research forward with the Long String Star V automation system.

Let’s Find You an Application That Helps Your Research
Get a call from your local Decode Science representative to help you find the best fit genomics products for you.
Or give us a call at: