Key Takeaways
Profiled 1 million human B cells in a single experiment
Detected 900,000+ unique paired clonotypes across Type 1 Diabetes, Multiple Sclerosis, Rheumatoid Arthritis, Crohn’s, Celiac, and healthy donors
Achieved sensitive detection of CDR3 regions, clonotypes, and full-length sequences at scale
Evercode BCR uncovered over 900,000 unique paired clonotypes across 24 samples in a single experiment. Negatively selected B cells from 12 healthy donors were purchased, while pan B cells were isolated from 12 autoimmune-diseased human PBMCs and fixed using the Evercode Cell Fixation Kit v3 to preserve cell structure and protect RNA integrity. Fixed samples were stored at -80°C until all were ready for combined processing with the Evercode BCR Mega Kit. Whole transcriptome and BCR-specific libraries were sequenced on the Illumina Novaseq X, and data were analyzed using Parse Biosciences’ Analysis Pipeline v1.3.0. Clustering with Seurat 5.0 showed that the majority of cells corresponded to major B cell subtypes, as illustrated in the UMAP below (Figure 1).
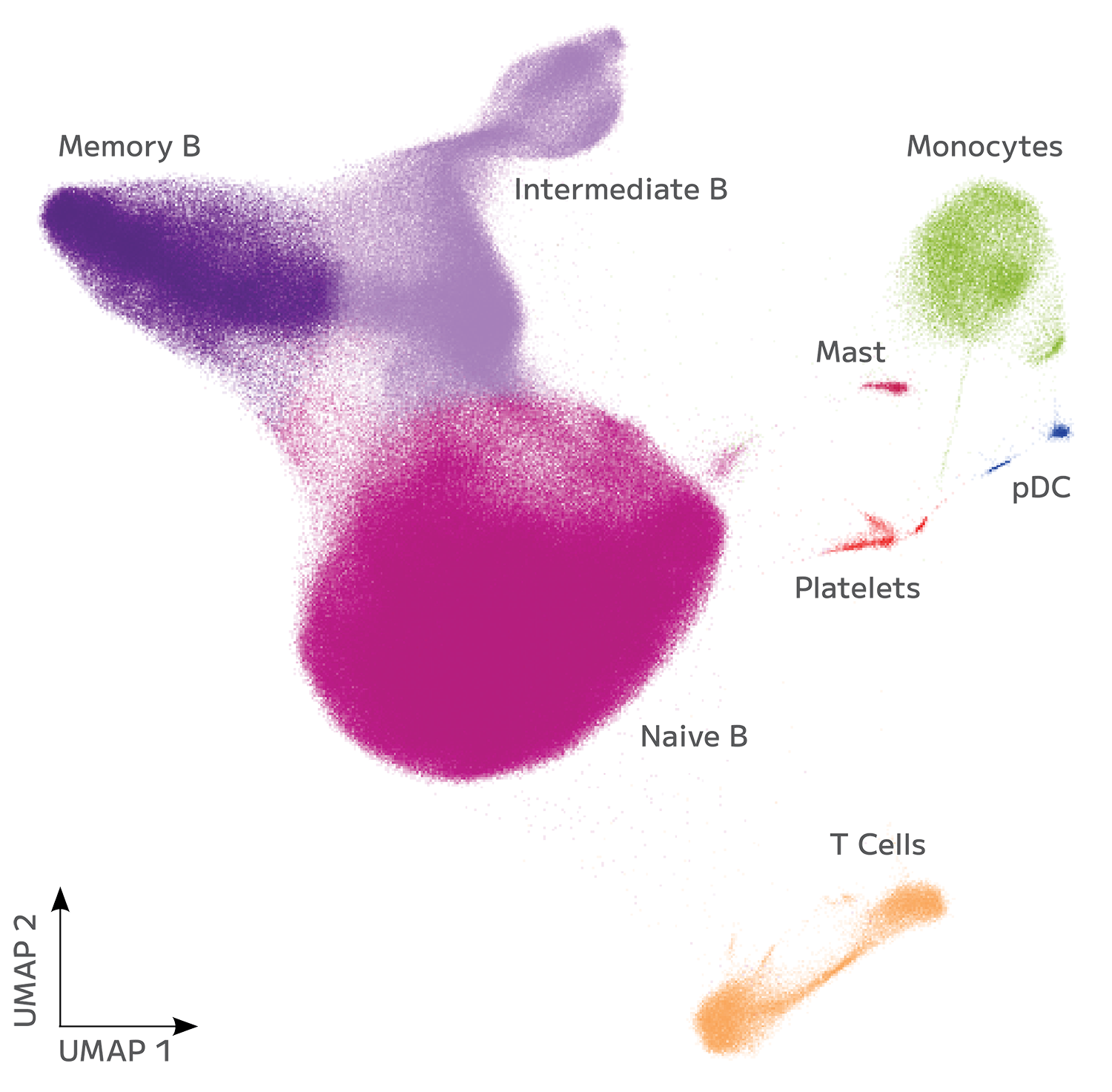
The assay demonstrated high sensitivity, detecting paired heavy and light chains in up to 89% of cells (Figure 2).


ANZ Market Manager - Research Genomics
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.