PRODUCTS
The utilization of single-site variant libraries is proving invaluable for researchers aiming to delve into a protein’s sequence space and understand the intricate relationship between sequence, protein structure, and function. This innovative approach allows for a comprehensive exploration of variants, providing crucial insights into the molecular dynamics of proteins.
The construction of Twist Site Saturation Variant Libraries takes protein engineering to the next level. Leveraging advanced massively parallel oligonucleotide synthesis through Twist’s proprietary silicon-based DNA synthesis platform, these libraries ensure a precise and controlled crafting of variants. With complete mastery over codon usage, high uniformity at each site, and rigorous quality control verified through next-generation sequencing (NGS), these libraries offer researchers the flexibility to conduct experiments with ease. Whether opting for one position per well in a 96-well plate or pooling all positions in a single tube, this methodology enables the screening of 1 to 20 different amino acids at each position, further enhancing the adaptability and efficiency of protein engineering studies.
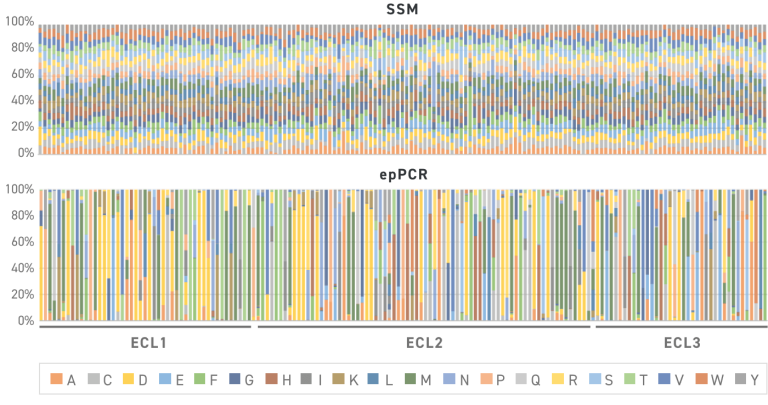
Site Saturation Libraries, particularly those constructed by Twist, revolutionize the exploration of sequence space in protein engineering by eliminating codon bias and preventing unwanted substitutions. In contrast to conventional methods like degenerate and NNK approaches, Twist Site Saturation Variant Libraries (SSVLs) offer a superior solution. Traditional methods, such as error-prone PCR and degenerate approaches, often suffer from poor repetitive yield, resulting in less than 50% full-length product in typical libraries. In comparison, Twist libraries yield more usable variants, significantly increasing the effective library size. The figure depicting the observed distribution of amino acids across 65 positions in a protein illustrates the efficiency of Twist SSVLs in maintaining the expected ratios of variants, showcasing their ability to provide highly uniform variant libraries.
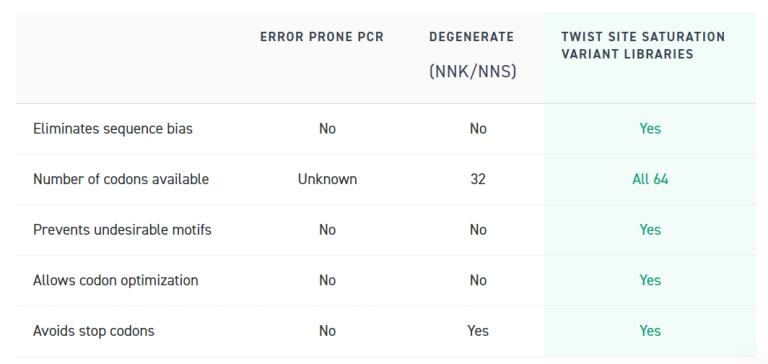
Comparing key features, Twist SSVLs outshine error-prone PCR and degenerate approaches on various fronts. They eliminate sequence bias, utilize all 64 codons, prevent undesirable motifs, allow codon optimization, and avoid stop codons. This comprehensive set of advantages positions Twist SSVLs as a powerful tool for efficient sampling of a protein’s sequence space in screening assays, demonstrating their superiority in precision, reliability, and overall effectiveness in protein engineering studies.
Get a call from your local Decode Science representative to help you find the best fit genomics products for you.
Or give us a call at:
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.