Download Instant Resources
MosaiX™ Library Prep Kit from seqWell combines the speed of tagmentation with the precision of ligation-based methods — without compromise. At its core is TnX, a next-generation engineered transposase that dramatically reduces the insertion site bias associated with conventional Tn5 enzymes. The result is libraries with exceptional molecular complexity, uniform coverage, and minimal duplication — from as little as 1 ng of input DNA.
Whether you’re scaling population genomics studies, performing whole genome or exome sequencing, or running targeted capture panels across human, plant, or animal samples, MosaiX delivers publication-ready data with a workflow that fits into a single morning. Directional tagmentation means you spend less time troubleshooting and more time generating insights.
Engineered for reduced insertion site bias compared to standard Tn5, TnX consistently accesses difficult genomic regions — including clinically relevant exome targets that other methods miss.
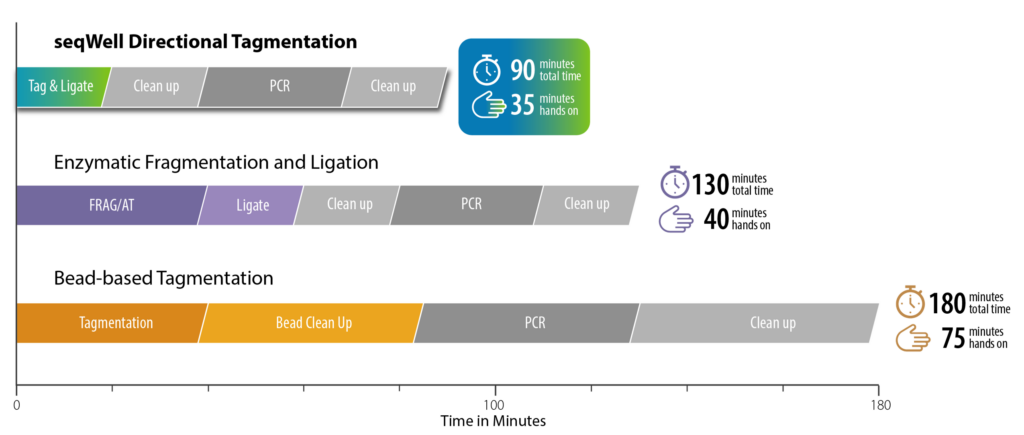
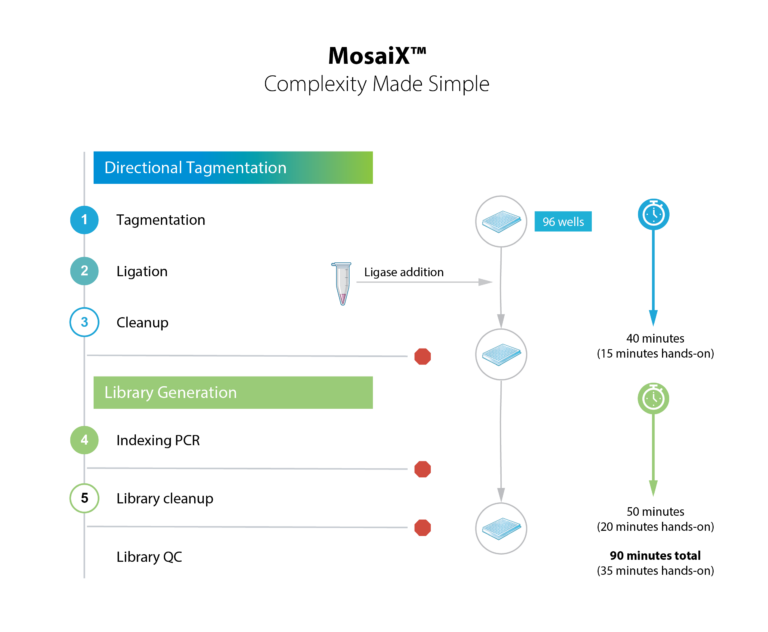
From DNA to sequencer-ready library in under two hours. Minimal hands-on steps mean you can process more samples with less effort and fewer errors.
MosaiX libraries routinely outperform bead-linked Tn5 preparations in library complexity and duplication rates — giving you more usable data per sequencing run.
Works with 1–50 ng gDNA in common buffers (Tris, TE, water). Compatible with all Illumina platforms, plus Element AVITI™ and Complete Genomics via conversion kits.
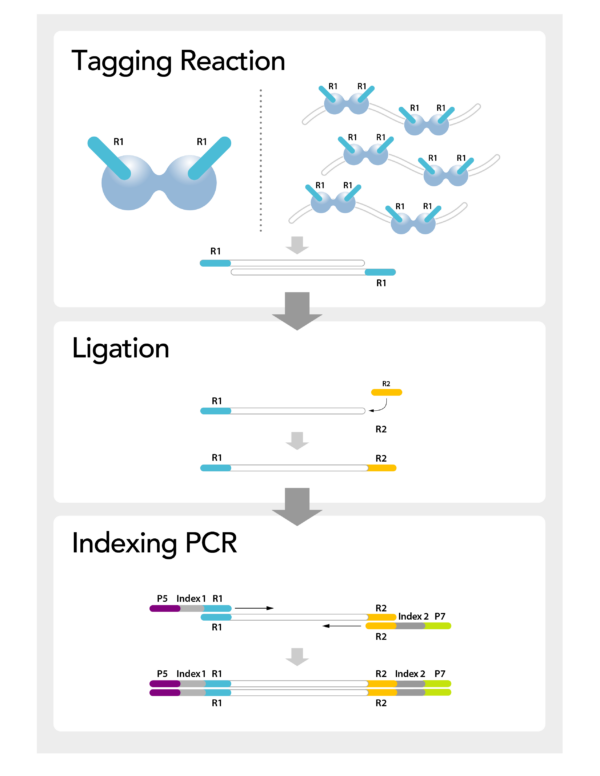

Clinical Sales Manager - ANZ Country Manager - NZ
Need help choosing the right kit for your application? Our technical specialists are ready to advise — reach out now and we’ll respond within the hour.
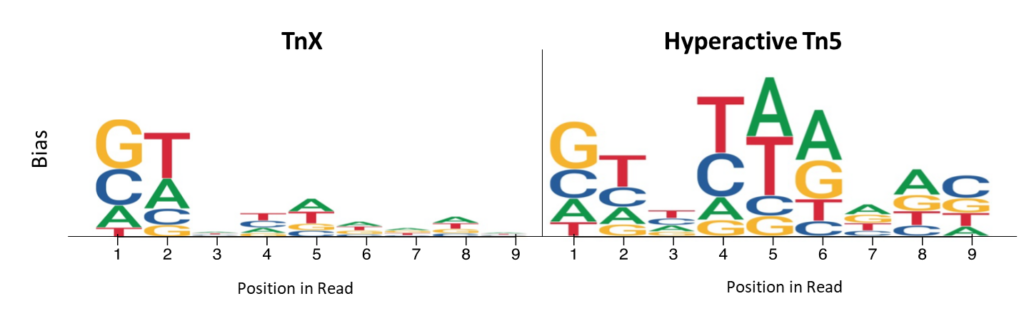
Read start site insertion bias was measured by examining the frequency of bases in the first 9 bases of each read. Positions with higher per-base nucleotide bias are represented by heights for hyperactive Tn5 and TnX, and illustrate the reduced bias of TnX.
comes with trade-offs: insertion bias, lower complexity, and missed targets. Ligation methods deliver quality but demand time and technical finesse. MosaiX bridges that gap.
every percentage point in duplication rate and every missed exon target translates to wasted sequencing spend and compromised variant calls. With MosaiX, you're not choosing between throughput and data quality — you're getting both.
achieve higher coverage uniformity and capture difficult genomic regions that bead-linked Tn5 preparations consistently miss. If your research depends on complete, unbiased representation of the genome, this is the kit that delivers.
Benchmark-Matched Quality With a Fraction of the Effort
When evaluated against the gold standard of enzymatic fragmentation followed by ligation, MosaiX-prepared libraries delivered virtually identical exome metrics at 6 Gb sequencing depth. But here’s where it gets interesting: compared to bead-linked Tn5 tagmentation, MosaiX consistently outperformed across the metrics that matter most — lower duplication rates, higher library complexity (as measured by HS Library Size), and fewer zero-coverage targets.
That last point deserves emphasis. Zero-coverage targets represent gaps in your data — regions you sequenced but couldn’t see. In exome studies, those gaps can mean missed variants in clinically actionable genes. MosaiX closes those gaps.
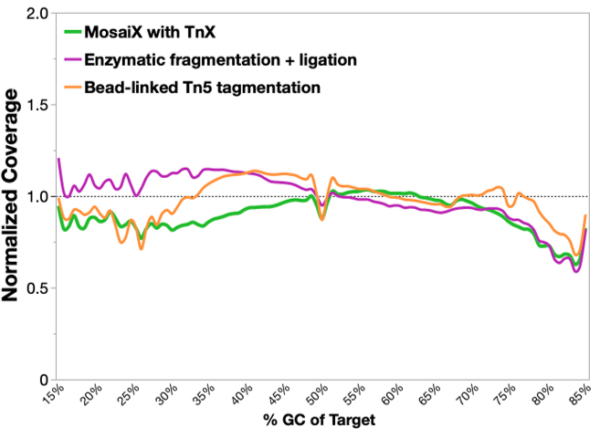
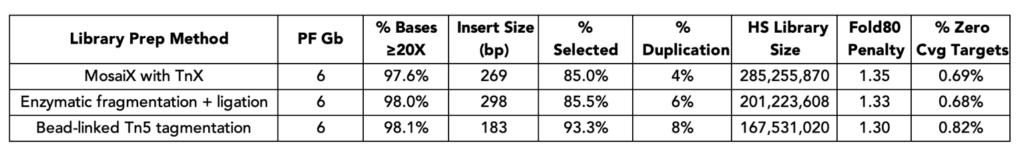
50 ng NA12878 genomic DNA (Genome in a Bottle reference) was used across all conditions. Libraries were prepared according to each manufacturer's protocol, captured using Twist Bioscience Exome 2.0 panel with standard workflow, and sequenced on NextSeq 2000. Data were down-sampled to 6 Gb per library and aligned to Twist exome capture targets on hg38.
Standard bead-linked tagmentation using conventional Tn5 has a known weakness: insertion site sequence bias. This bias creates systematic blind spots — regions of the genome where the transposase preferentially avoids inserting, resulting in poor or absent coverage.
In exome sequencing, this isn’t a minor inconvenience. It means clinically relevant targets can fall into coverage gaps, leading to missed variant calls in genes that could inform diagnosis or treatment decisions.
TnX was engineered specifically to address this limitation. Its reduced insertion bias, combined with the higher molecular complexity of MosaiX libraries, enables access to difficult genomic regions that Tn5-based methods routinely underrepresent.
The practical outcome: fewer zero-coverage targets, more complete exome representation, and greater confidence in your variant calls.
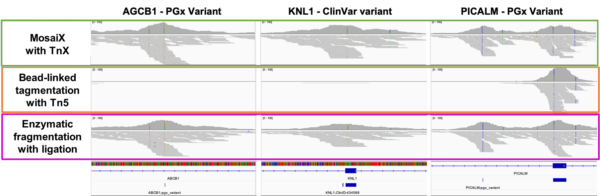
At matched sequencing depth (105 Gb, down-sampled from NovaSeq X+ 25B), MosaiX libraries achieved higher mean coverage than bead-linked Tn5 preparations. Duplication rates were lower. Estimated library size — a direct indicator of molecular complexity — was higher.
What does this mean in practice?
You’re extracting more unique, mappable information from every gigabase of sequencing output. For population-scale studies or projects where sequencing cost is a limiting factor, that efficiency translates directly to better data economics.
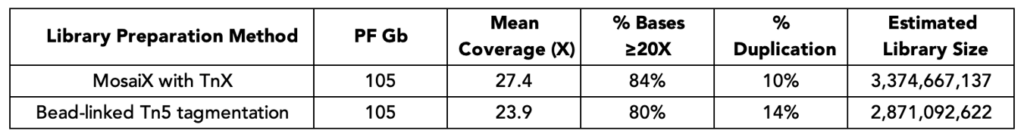
Method: 50 ng of NA12878 DNA (Genome in a Bottle) was used in both and libraries were prepared following manufacturers’ user guides. Sequencing was performed on a lane of a NovaSeq X+ 25B flow cell, down-sampled to 105 Gb each, then aligned to hg38.

Clinical Sales Manager - ANZ Country Manager - NZ
Ready to trial MosaiX in your lab?
Get in touch with our team — we’ll have pricing and availability to you within 24 hours.

TnX Read 1 Tagging Reagent
5X Reaction Buffer
Tagmentation Enhancer
Read 2 Adapter
DNA Ligase
2X Amplification Ready Mix
MAGwise Paramagnetic Beads
Diluent
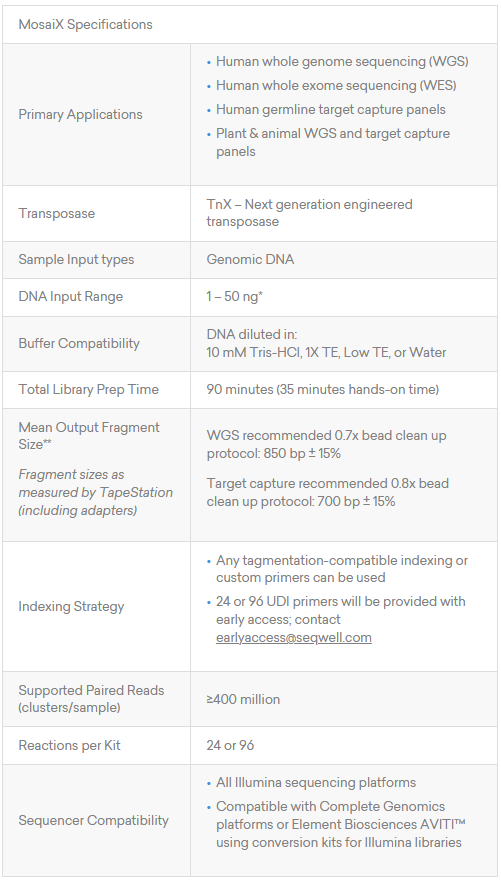
MosaiX is optimised for purified genomic DNA from human, plant, or animal sources. Input can range from 1–50 ng, though inputs below 5 ng may require optimisation of adapter concentration and PCR cycles.
Yes. MosaiX libraries are compatible with all Illumina sequencing platforms. For Element AVITI™ or Complete Genomics systems, use the appropriate Illumina library conversion kit.
TnX is an engineered transposase with significantly reduced insertion site sequence bias. This results in higher library complexity, lower duplication rates, and better access to difficult genomic regions compared to conventional Tn5-based methods.
MosaiX is compatible with any tagmentation-compatible indexing primers. Kits include 24 or 96 unique dual index (UDI) primers.
Absolutely. MosaiX has been validated for whole exome sequencing (WES) and targeted capture panels, with benchmarking data showing improved performance over bead-linked Tn5 methods for these applications.
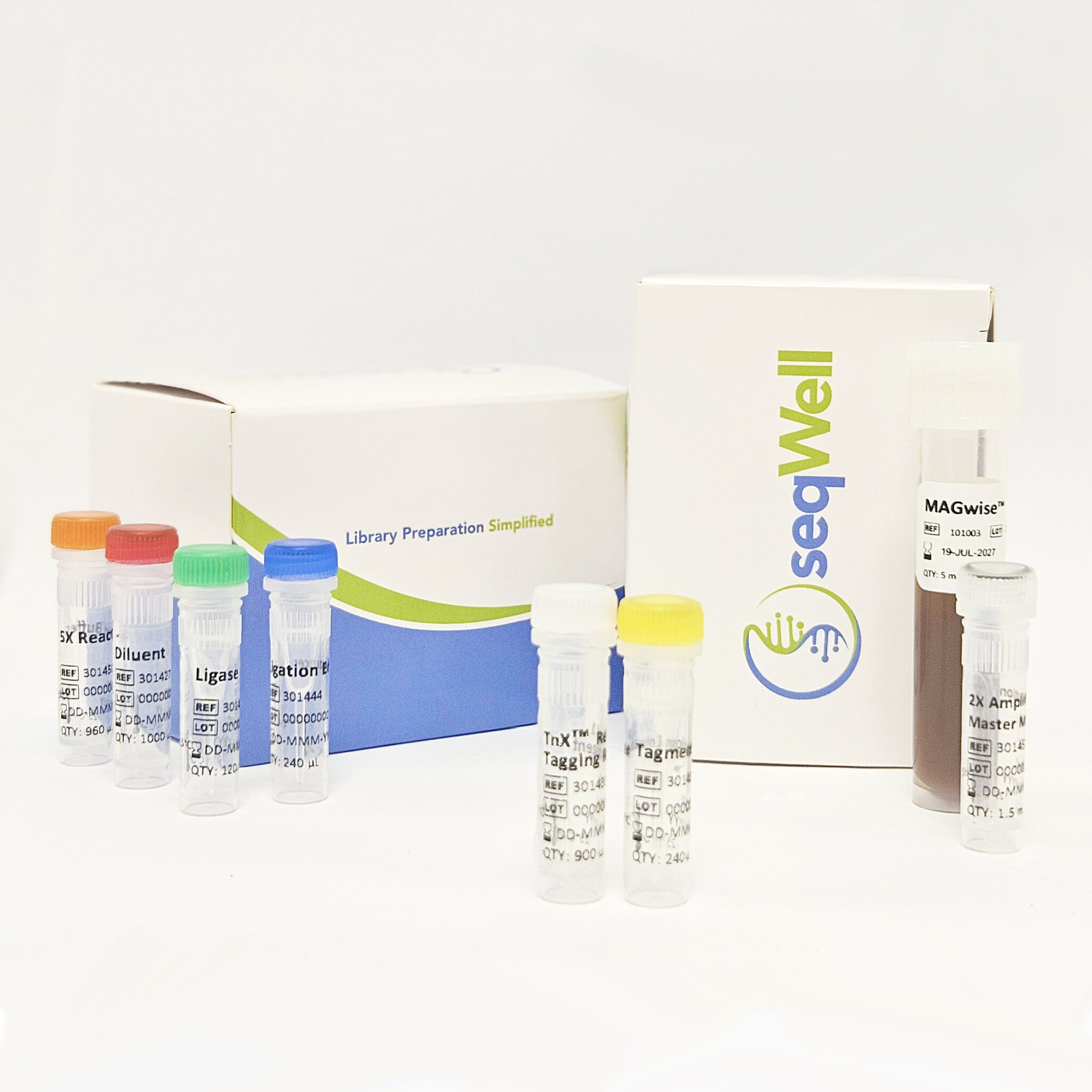
Each kit contains: TnX Read 1 Tagging Reagent, 5X Reaction Buffer, Tagmentation Enhancer, Read 2 Adapter, DNA Ligase, 2X Amplification Ready Mix, MAGwise Paramagnetic Beads, and Diluent.
We only need below information to serve you better. Decode Science respects your privacy and will never spam you with unrelated content.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.