
The Twist TrueAmp Library Preparation Kit is a precision-engineered solution for next-generation sequencing target enrichment workflows, purpose-built for laboratories that cannot afford to lose data from difficult samples. At its core is the Twist TrueAmp polymerase — a high-fidelity amplification enzyme optimised to suppress amplification-induced errors and GC-related bias, delivering the coverage uniformity your downstream variant analysis demands. Whether you’re working with pristine input or fighting the constraints of degraded FFPE material, TrueAmp is designed to keep your libraries representative and your results reproducible.
Paired with optimised enzymatic fragmentation and a high-efficiency ligation step, the TrueAmp workflow delivers tunable, consistent insert sizes across a wide range of input quantities — making it well-suited to mixed-input workflows and scaled sequencing operations. From pre-capture yield to on-target coverage, every metric reflects the same design intent: higher library complexity, fewer reruns, and greater confidence in variant calls from the first pass. As your authorised Australian distributor, Decode Science can advise on kit configurations, protocol optimisation, and compatibility with your current target enrichment panels.


Exclusive Offer for now!!
50% off 16 sample workflow kits & 50% off 96 sample kits.
When working with low-input or degraded FFPE DNA, pre-capture library yield is the first indicator of whether a prep has succeeded or failed. TrueAmp consistently delivers higher pre-capture yields alongside stronger mean target coverage — without sacrificing library complexity. The result is more sequenceable molecules from the same difficult starting material, fewer failed runs, and less pressure on irreplaceable samples.
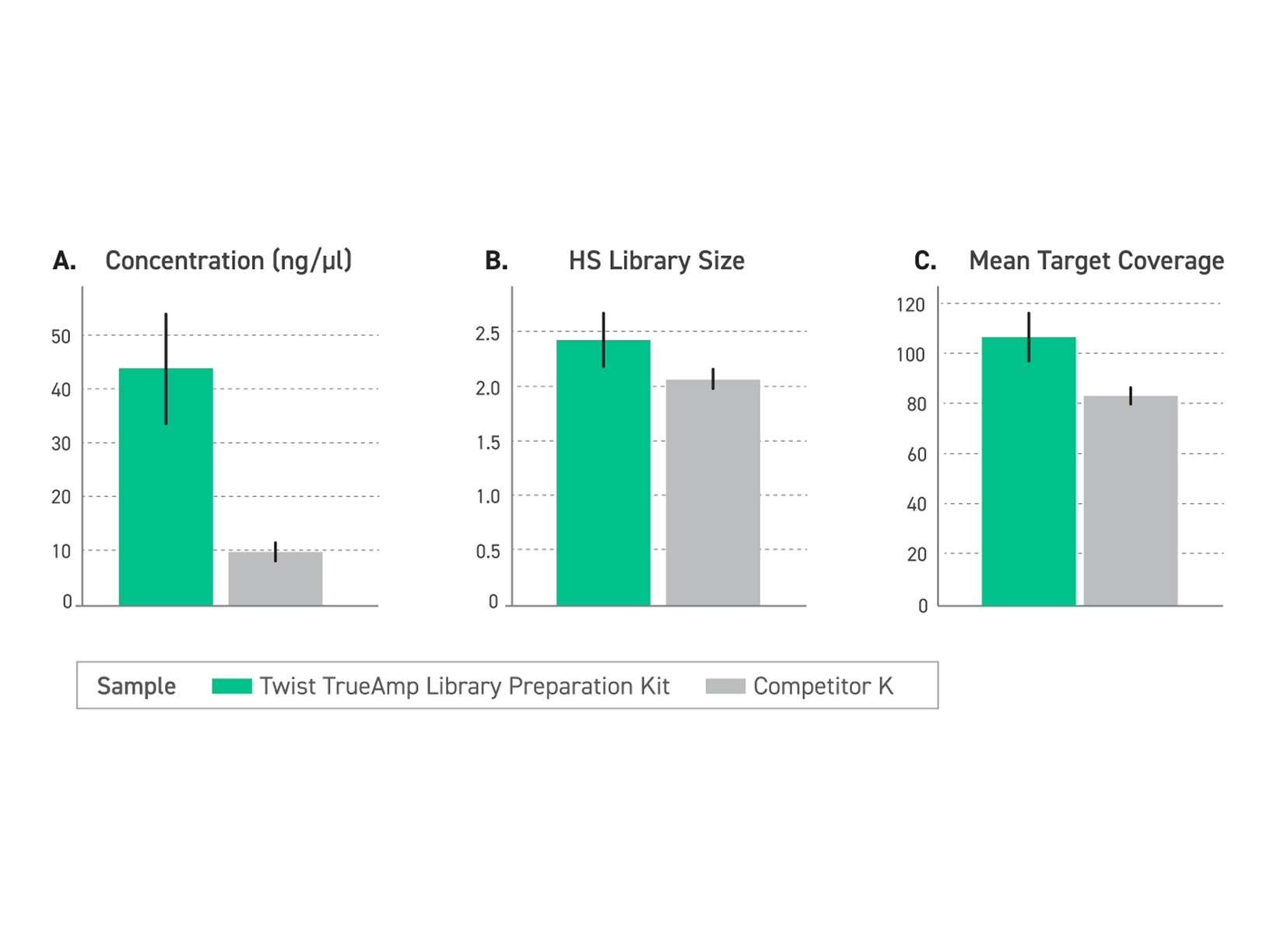
Figure 1. Performance comparison of enriched libraries with low-input FFPE degraded samples (DIN <2.2) between TrueAmp Library Preparation Kit and competitor K kit, demonstrating the optimal solution for challenging sample applications. (A) The Twist TrueAmp Library Preparation Kit generates superior pre-capture library yield, indicative of high library construction and amplification efficiency. (B) The Twist TrueAmp Library Preparation Kit shows higher library complexity when compared with the competitor’s kits. This allows for more unique DNA molecules that are sequenceable in the library, reducing sequencing costs. (C) Achieves higher coverage.
Gaps in coverage are not just an inconvenience — in target enrichment workflows, they represent missed variants and incomplete answers. TrueAmp reduces the proportion of zero-coverage targets across captured regions, improving coverage uniformity and giving you greater confidence that every target in your panel has been adequately interrogated. What you sequence is what you intended to sequence.
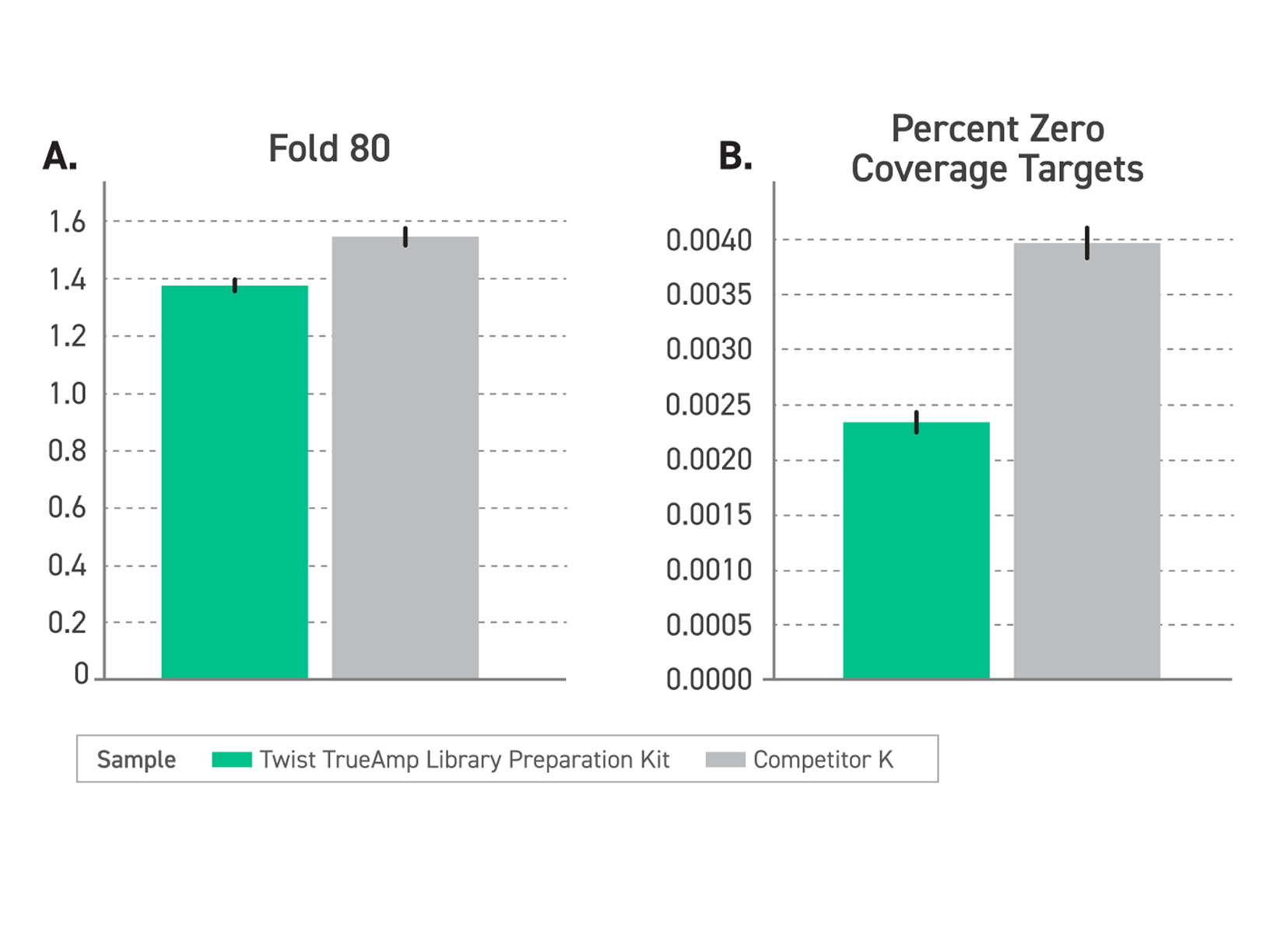
Figure 2. Performance comparison of enriched libraries with low-input FFPE degraded samples (DIN <2.2) between TrueAmp Library Preparation Kit and competitor K kit, demonstrating the optimal solution for challenging sample applications. (A) Delivers excellent coverage uniformity, measured by a lower fold-80 base penalty. (B) Reduced regions with no coverage, measured by Percentage of Zero Coverage Targets.
GC content remains one of the most persistent sources of coverage bias in NGS library preparation. The TrueAmp Polymerase Mix is specifically formulated to maintain uniform amplification across both high- and low-GC regions — and critically, this performance holds even as PCR cycle number increases. For panels that include regulatory elements, repetitive regions, or GC-skewed targets, this translates directly to more complete and trustworthy data.
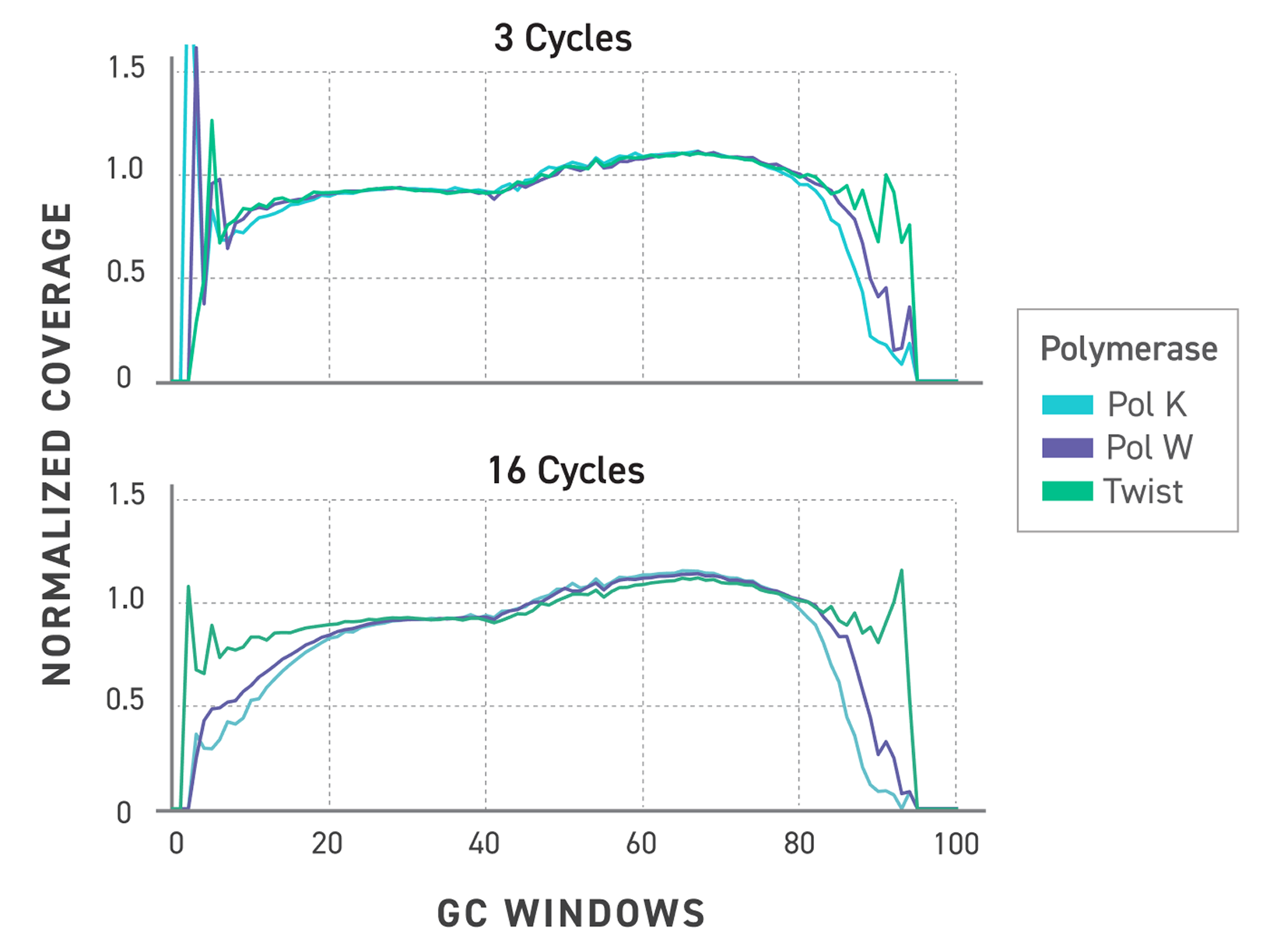
Figure 3. Normalized GC bias trace showing improved coverage of the Twist TrueAmp polymerase. Libraries were prepared with Twist TrueAmp Library Preparation Kit and amplified with different polymerases and cycles.
Upper panel: Normalized coverage against GC window plots comparing polymerases at 3 cycles of PCR.
Lower panel: Normalized coverage against GC window plots comparing polymerases at 16 cycles of PCR.
Fragment size consistency underpins downstream QC, sequencing performance, and batch-to-batch reproducibility. TrueAmp delivers tightly controlled insert size distributions across a broad range of input DNA quantities, making it well-suited to mixed-input batching strategies and high-throughput workflows where uniformity at scale is non-negotiable.
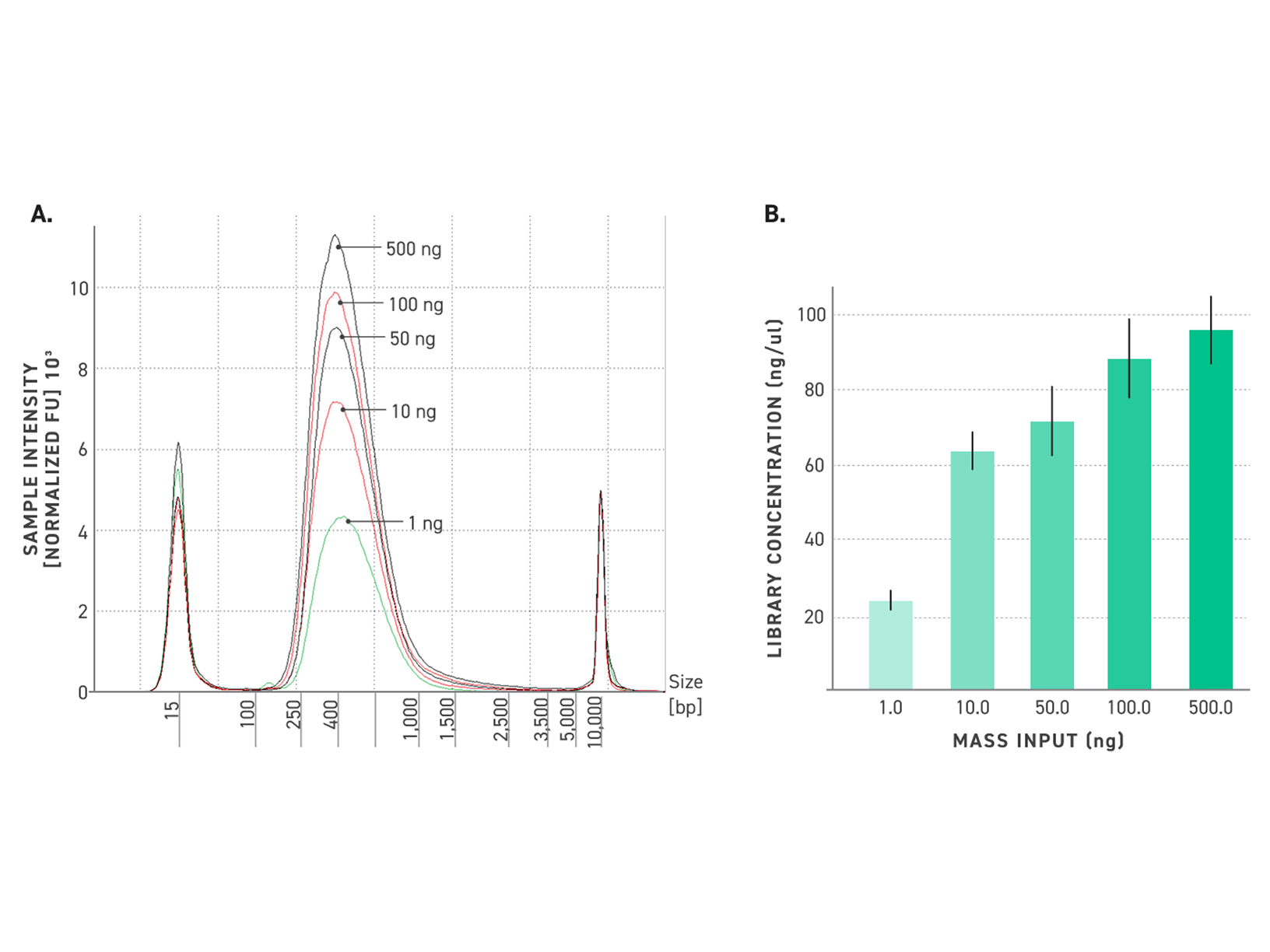
Figure 4. Reliable library size with Twist TrueAmp Library Prep Kit, even from ultra-low inputs.
500 ng, 100 ng, 50 ng, 10 ng and 1 ng (gDNA) were fragmented at 32°C. 3, 5, 6, 8, 10, and 14 cycles of PCR were utilized for amplification, respectively. Samples have been performed in duplicates.
A: Electropherograms of NGS libraries generated with the Twist TrueAmp Library Preparation Kit.
B: Concentration of libraries after amplification for various DNA inputs.
Not every workflow demands the same library architecture. TrueAmp’s enzymatic fragmentation step produces repeatable, tunable insert sizes that can be adjusted to match your sequencing platform requirements and downstream analysis needs — providing flexibility without sacrificing the reproducibility your pipeline depends on.
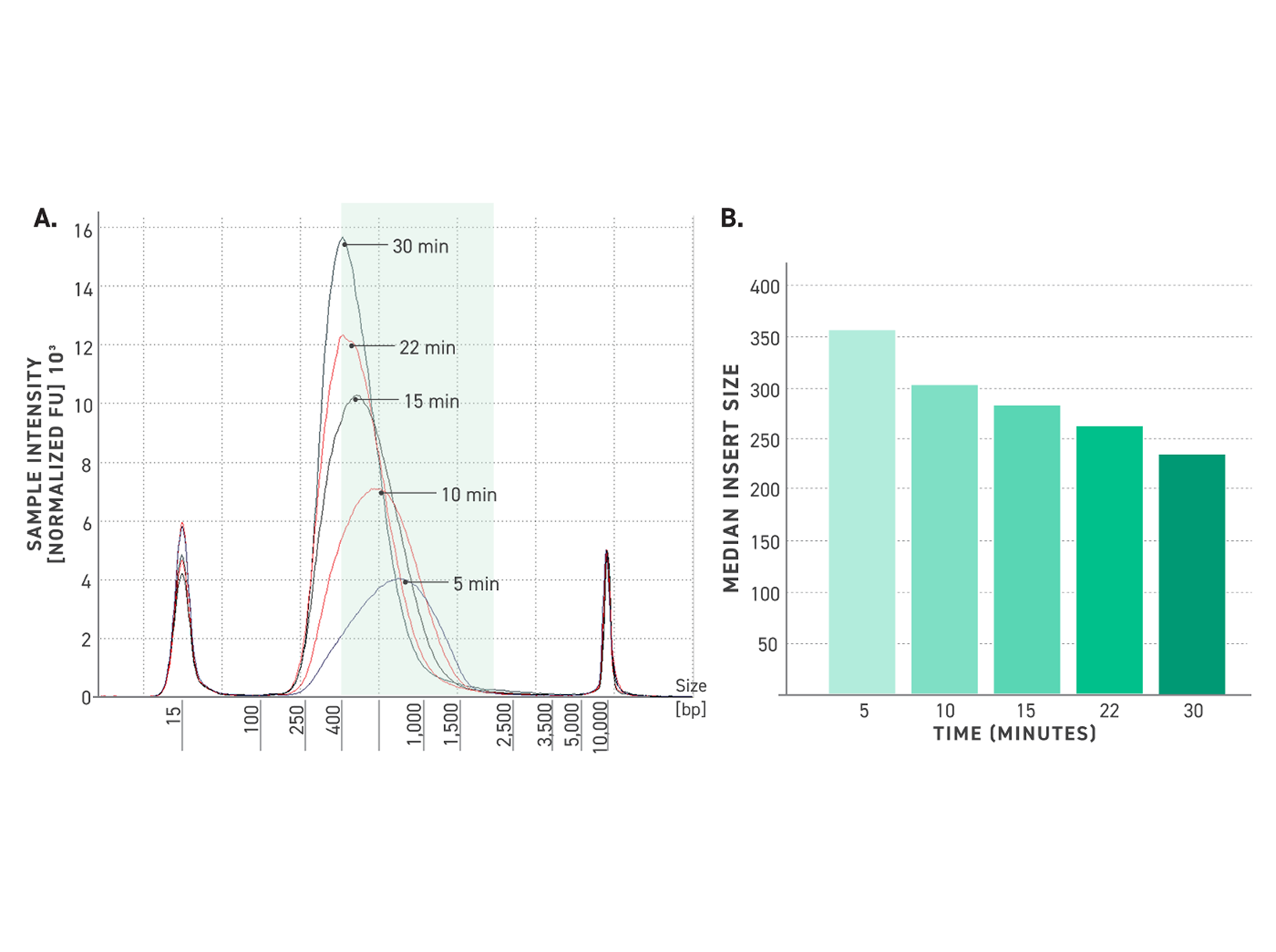
Figure 5. Tunability of Twist TrueAmp Library Prep Kit.
A: Five electropherograms of NGS libraries generated using differing fragmentation times. 50 ng of high-quality gDNA was fragmented for various times at 32°C. 6 cycles of PCR were utilized for amplification.
B: Median insert size vs time. 50 ng of high-quality gDNA was fragmented for various times at 32°C. Amplification was performed using 6 cycles of PCR. Samples were captured using the Twist Exome 2.0 panel.
Three steps. Consistent results.
The TrueAmp workflow is built around three precision-optimised steps that together deliver libraries you can sequence with confidence:
Step 1 — Enzymatic Fragmentation: Extracted DNA is fragmented enzymatically to produce consistent, tunable insert sizes — no sonication required, no shear-related variability.
Step 2 — Adapter Ligation: An optimised Twist ligase formulation maximises adapter conversion efficiency while minimising ligation bias, preserving molecular diversity from the very first step.
Step 3 — Amplification via TrueAmp Polymerase: High-fidelity amplification boosts yield from challenging templates, maintains coverage uniformity across GC extremes, and supports sensitive downstream variant detection.
In translational research and clinical genomics, the samples most critical to a study are often the ones that are hardest to sequence. Archival FFPE blocks, fine-needle aspirates, liquid biopsy specimens, and low-cellularity tumour sections are routinely degraded, limited in quantity, or variable in quality — and standard library prep kits frequently fail them.
TrueAmp was engineered specifically for this challenge. Its impact is most pronounced where it matters most:
Low VAF variants in heterogeneous tumour samples require error suppression and high complexity to call reliably. TrueAmp’s fidelity advantage directly supports this.
Legacy tissue samples carry irreplaceable longitudinal or retrospective data. Higher pre-capture yields from degraded input means more of that data becomes sequenceable.
Coverage uniformity across all captured targets — including GC-extreme regions — is the difference between a complete and an incomplete picture of your panel of interest.
Reproducible fragment sizes and consistent performance across input ranges simplify batch QC, reduce failed libraries, and increase instrument utilisation.

Clinical Genomics Manager - ANZ & Country Manager - NZ
Bias-free whole genome libraries from high-quality input DNA, no amplification required
Design and order target enrichment panels tailored to your gene list or genomic region of interest
Comprehensive exome capture panel with proven uniformity across canonical and difficult targets
TrueAmp is specifically validated for challenging input types including FFPE-derived DNA (DIN <2.2), low-input samples, and variable-quality clinical specimens. It also performs well with high-quality genomic DNA across a wide input range.
Input requirements vary by sample type and downstream application. TrueAmp is designed to deliver high conversion efficiency even from low-input and degraded samples — contact Decode Science for guidance specific to your sample type and target enrichment panel.
Yes. TrueAmp is validated for use with both the Twist Universal Adapter System and the Twist UMI Adapter System, supporting error correction workflows for somatic variant detection and liquid biopsy applications.
Absolutely. TrueAmp is designed to integrate with the full Twist target enrichment ecosystem, including Twist Custom NGS Panels and standard off-the-shelf panels such as Twist Exome 2.0.
In head-to-head benchmarking against a leading competitor kit (Competitor K), TrueAmp generated superior pre-capture library yields, higher library complexity, and greater mean target coverage from the same degraded FFPE input — preserving more sequenceable molecules from precious, limited-quantity samples.
TrueAmp libraries are compatible with Illumina sequencing platforms. For platform-specific guidance, contact our team.
Protocols for TrueAmp with both the Universal Adapter System and UMI Adapter System are available from Decode Science on request.
We only need these information to serve you better. Decode Science respects your privacy and will never spam you with unrelated content.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.