
The Twist PCR-Free WGS Library Preparation Kit is engineered for researchers who need their sequencing data to reflect the genome as it actually exists — not as amplification has reshaped it. By eliminating the PCR amplification step entirely, this kit preserves native genome representation from the outset, removing the single greatest source of coverage bias in whole genome sequencing workflows. Built on Twist’s optimised enzymatic fragmentation and high-efficiency ligation chemistry, the workflow delivers consistent insert sizes, minimised ligation bias, and uniform genome-wide coverage that supports confident variant characterisation from the first run.
Designed to scale from individual research projects to population-level sequencing studies, the PCR-Free WGS kit supports multiplexing of up to 1,536 samples per run via Twist’s full-length unique dual index (UDI) adapters — making it equally suited to high-throughput core laboratories and discovery programmes with large cohort requirements. As your authorised Australian distributor, Decode Science can support you with kit configurations, protocol guidance, and compatibility assessment for your sequencing platform and informatics pipeline.


Exclusive Offer for now!!
50% off 16 sample workflow kits & 50% off 96 sample kits.
The PCR-Free WGS kit delivers strong library yields and consistent library size distribution even as DNA input is reduced, outperforming competitor PCR-free workflows at equivalent input levels. Efficient ligation chemistry drives conversion across input amounts — producing sequencer-ready libraries that don’t require amplification rescue when input is reduced.
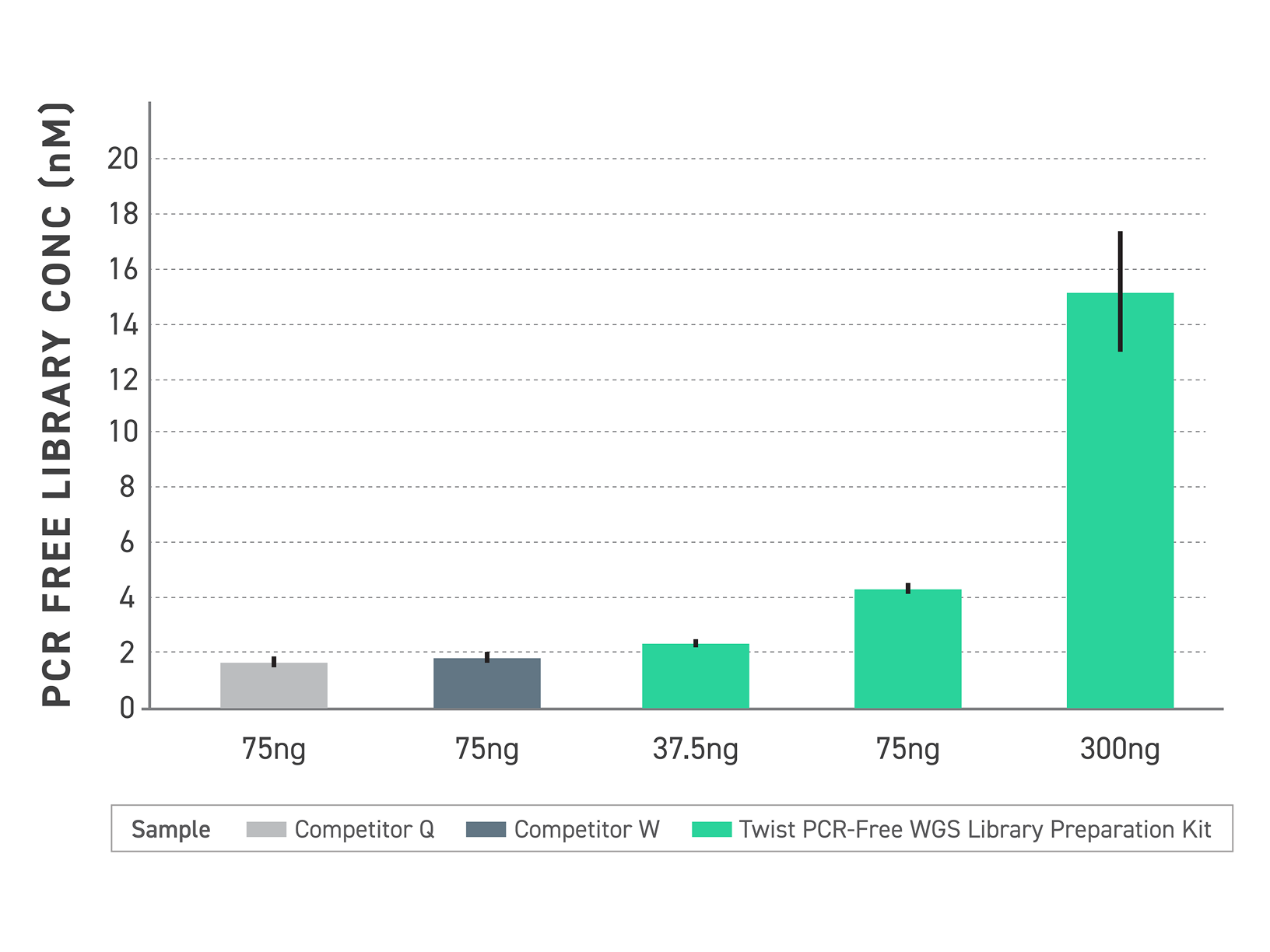
Figure 1. PCR-free library yield comparison across inputs. Final library concentration (nM) was measured following PCR-free library preparation using the indicated DNA inputs. Twist PCR-Free WGS Library Preparation generated higher library yields than competitor workflows at equivalent inputs and maintained strong performance at lower inputs.
Median insert sizes remain consistent across the full supported input range, from 300 ng down to 37.5 ng. Minimal read overlap across samples in this range confirms that fragment size control is maintained regardless of input quantity — supporting dependable genome-wide coverage without depth fluctuation between samples.
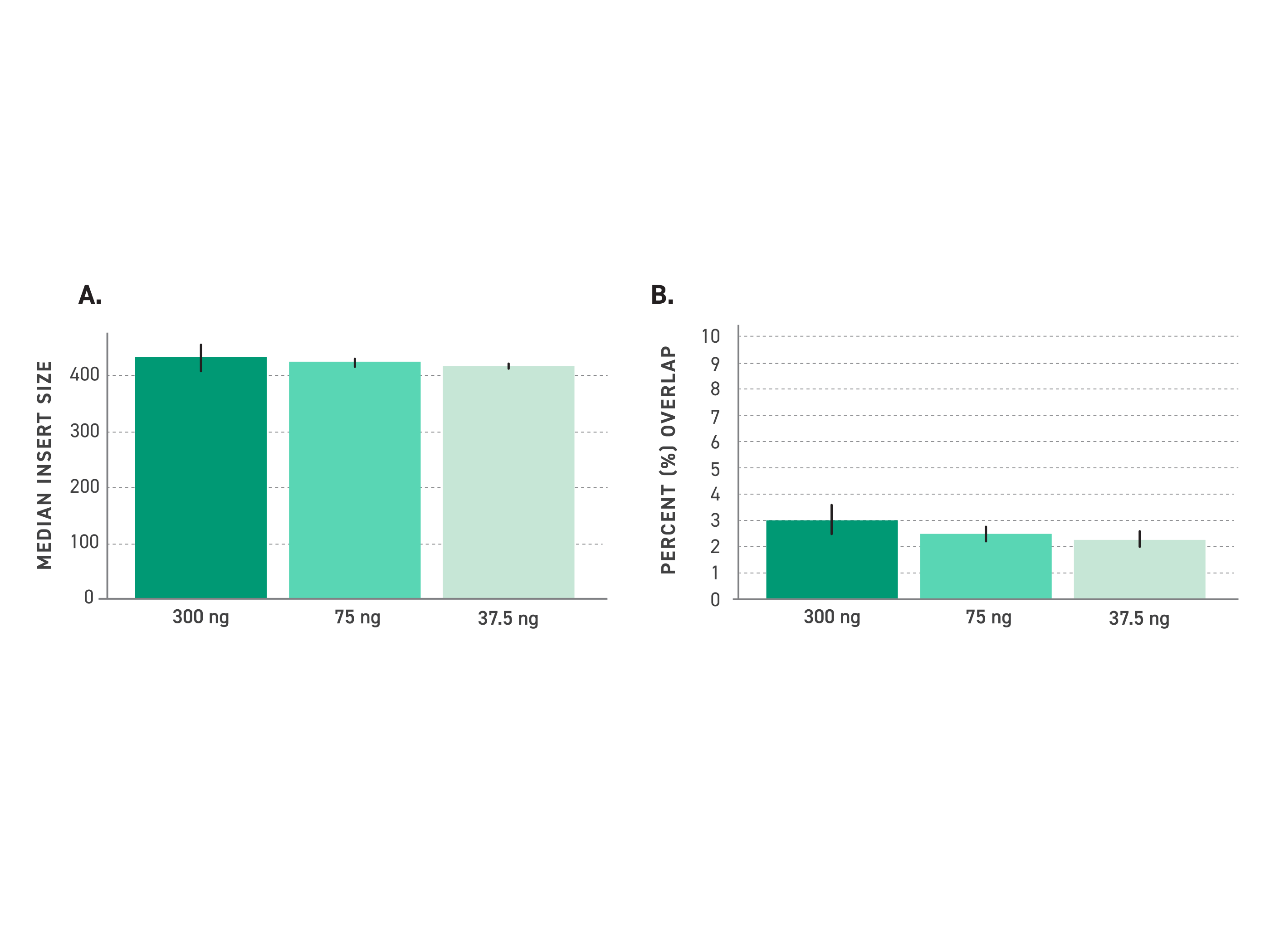
Figure 2. The kit produces consistently large inserts across a wide range of DNA samples. (A)Median Insert Size is represented for 300 ng, 75 ng, and 37.5 ng of NA12878 DNA sample input. (B) Percent Overlap (which measures how much paired-end sequencing reads redundantly cover the same DNA bases) is represented for 300 ng, 75 ng, and 37.5 ng of NA12878 sample input.
Enzymatic fragmentation parameters can be adjusted to produce insert size distributions matched to the requirements of your sequencing platform and read length. This gives laboratories flexibility to optimise the prep for their specific instruments and analysis pipelines without sacrificing inter-batch reproducibility.
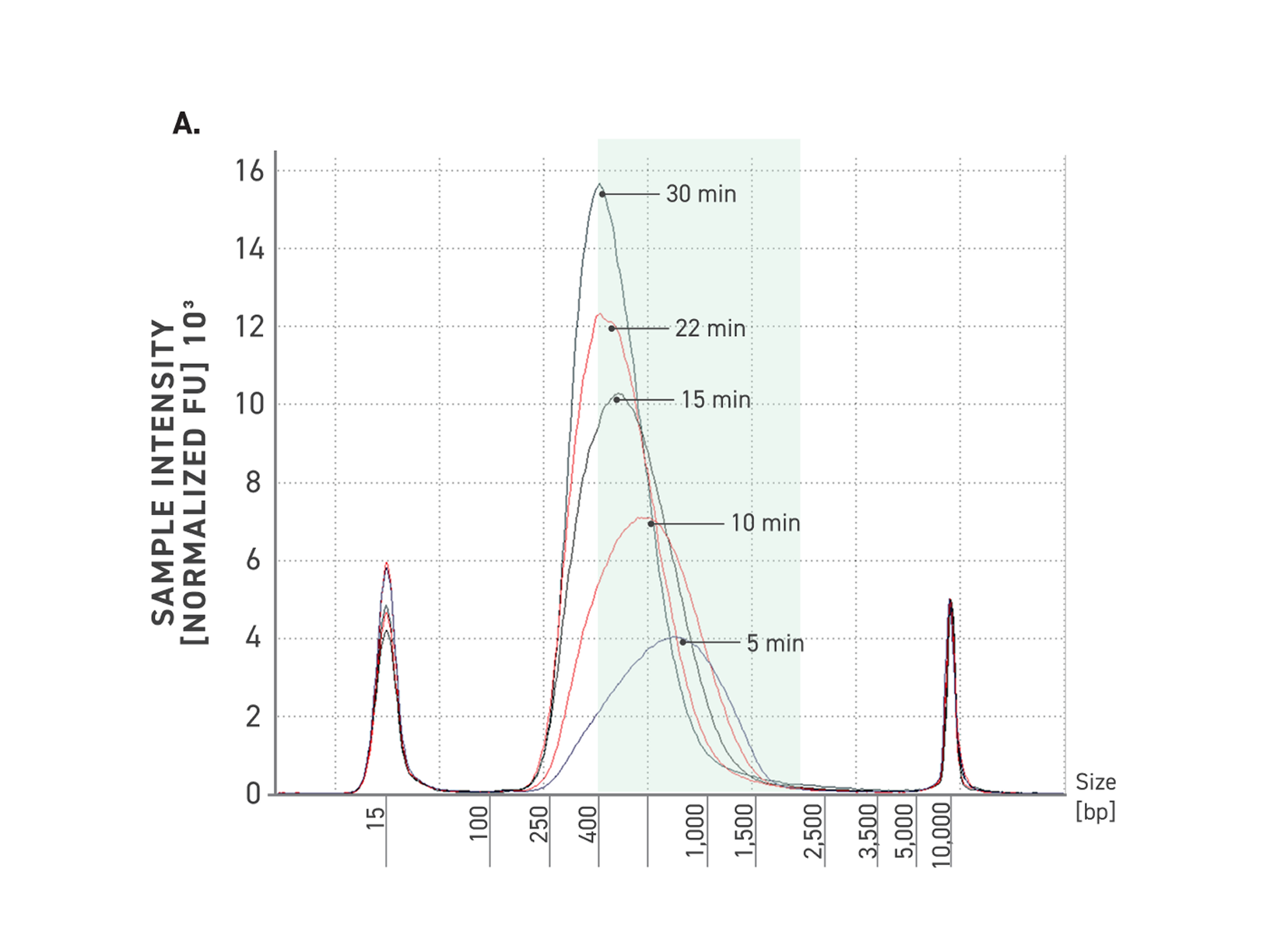
Figure 3. Tunability of Twist PCR-Free WGS Library Prep Kit.
A: Five electropherograms of NGS libraries generated using differing fragmentation times. 50 ng of high-quality gDNA was fragmented for various times at 32°C.
One of the most persistent sources of WGS data quality problems is differential coverage across GC content — PCR amplification exacerbates this by further enriching already-accessible fragments. By removing amplification, the PCR-Free WGS kit maintains even coverage across GC content categories regardless of input, yielding cleaner, more interpretable genome-wide data.
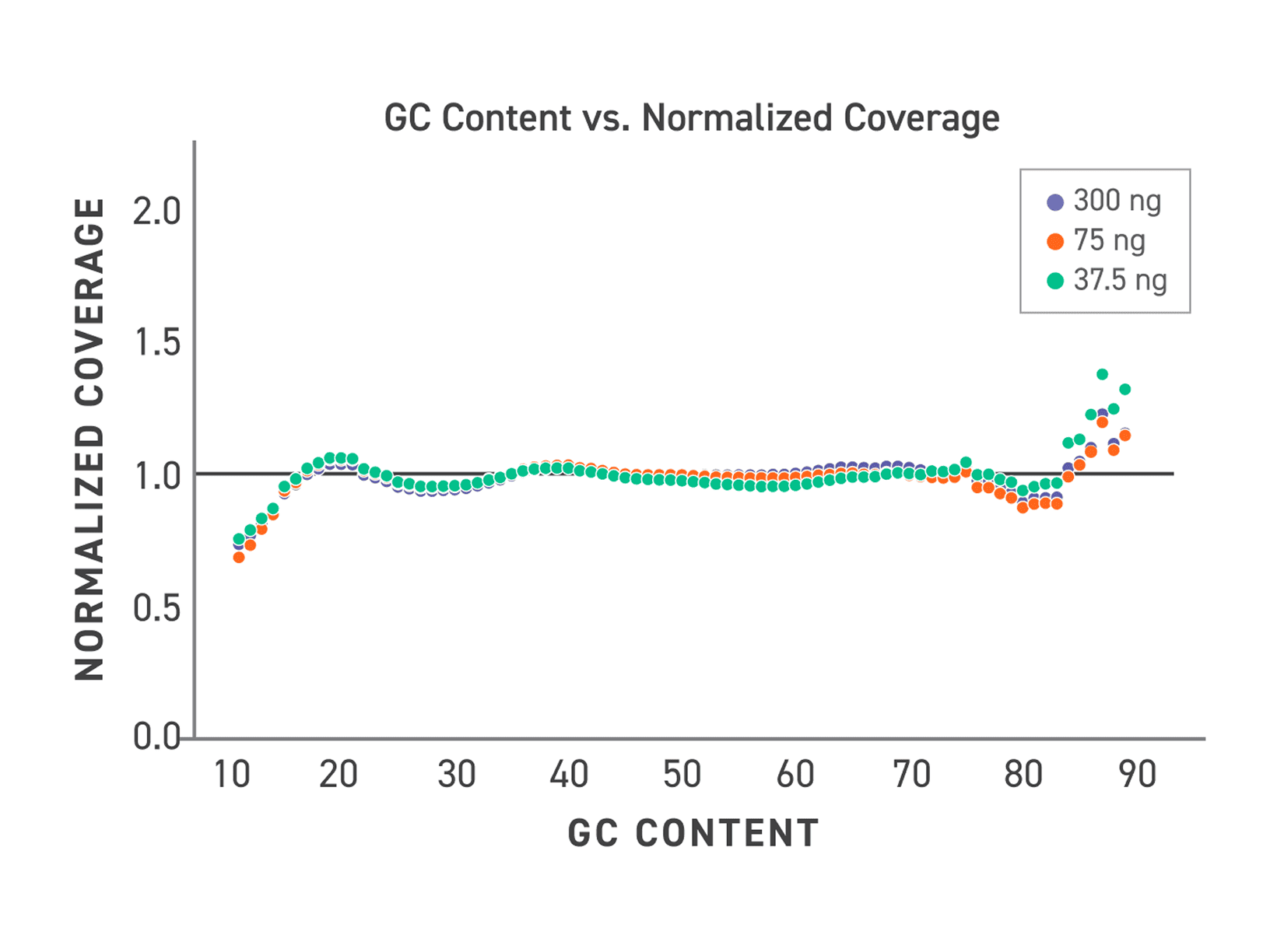
Figure 4. The Twist PCR-Free WGS Library Preparation Kit provides more consistent coverage distribution across varying GC content. The kit shows minimized GC bias even at low inputs, thereby reducing the need for additional sequencing to achieve uniform coverage and better detection of variants in the high- and low-GC content regions.
Two steps to sequencer-ready whole genome libraries.
The PCR-Free WGS workflow is deliberately streamlined — complexity is removed at the chemistry level, not pushed onto the operator.
Step 1 — Enzymatic Fragmentation: Input DNA is enzymatically fragmented to produce consistent, tunable insert sizes across all samples in a batch. No sonication, no shear-related variability — just controlled, reproducible fragmentation that sets the foundation for uniform downstream coverage.
Step 2 — Adapter Ligation: Twist’s optimised ligase chemistry maximises adapter conversion efficiency while minimising ligation bias. The result is a sequencer-ready library that accurately represents the molecular diversity of your input, with no amplification step introducing artificial enrichment of any genomic region.
Whole genome sequencing is increasingly the method of choice for variant discovery, structural analysis, and population-scale genomics — but the value of WGS data depends entirely on whether the library faithfully represents the genome that went into it. Amplification-based workflows introduce systematic biases that are difficult to distinguish from true biological signal, particularly in AT-rich regions, repetitive elements, and low-complexity sequences.
The PCR-Free WGS kit addresses this directly. Its impact is most relevant for:
Large-scale studies demand per-sample consistency, reproducible coverage distributions, and the ability to scale indexing without index hopping or sample cross-contamination. UDI adapter compatibility and 1,536-plex capacity make this feasible at production scale.
Accurate copy number and structural variant calls depend on even baseline coverage across the genome. Amplification-induced regional bias creates false signals that complicate interpretation; PCR-free prep removes this confounder at source.
Comprehensive, unbiased genome representation is the baseline requirement for germline variant calling, particularly in regions with extreme GC content or repetitive architecture that amplification-based methods handle poorly.
Consistent insert sizes, robust yields across a range of inputs, and multiplexing capacity that scales to 1,536 samples translate directly to higher instrument utilisation and lower per-sample sequencing cost.

Clinical Genomics Manager - ANZ & Country Manager - NZ
High-fidelity amplification-based library prep for target enrichment and challenging low-input or FFPE samples
Design and order target enrichment panels tailored to your gene list or genomic region of interest
Comprehensive exome capture panel with proven uniformity across canonical and difficult targets
Amplification-free workflows are more sensitive to input DNA quality than PCR-based methods, as there is no amplification step to recover yield from degraded material. High molecular weight, intact genomic DNA is recommended. If your samples are degraded or of variable quality, the Twist TrueAmp Library Preparation Kit may be a better fit — contact Decode Science to discuss.
The PCR-Free WGS kit maintains robust performance from 300 ng down to 37.5 ng DNA input, with consistent insert sizes and sequencing-ready yields across this range.
The kit is compatible with Twist’s full-length UDI adapter system, supporting multiplexing of up to 1,536 uniquely indexed samples per sequencing run.
In direct comparisons, the PCR-Free WGS kit delivers higher library yields than competitor PCR-free workflows at equivalent inputs. The absence of amplification removes duplicate reads from the sequencing output, meaning a greater proportion of reads generated are informative — improving effective sequencing depth per run.
The kit is compatible with Illumina sequencing platforms. Contact Decode Science for guidance on read length optimisation and coverage depth requirements for your specific platform and study design.
Yes. Amplification-free library prep is particularly well-suited to structural variant and copy number variant analysis, where even baseline coverage is essential. Removing PCR-induced regional enrichment reduces false positive signals in copy number calling.
The Twist PCR-Free WGS Library Preparation Kit protocol is available from Decode Science on request.
We only need these information to serve you better. Decode Science respects your privacy and will never spam you with unrelated content.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.