Detect pathogenic variants, copy number changes, and repeat expansions with clinically validated methods — enabling informed reproductive decisions and genetic counseling.
Traditional short-read NGS struggles with genes like SMN1 and FMR1 due to high homology or repeat expansions.
Mis-mapping or inaccurate repeat counts can lead to inconclusive or misleading results, delaying patient decisions.
CFTR: Only pathogenic variants tested, reducing the risk of reporting benign variants as carriers.
SMN1: Detects complex carrier states, including 2+0 carriers, which are missed by standard assays.
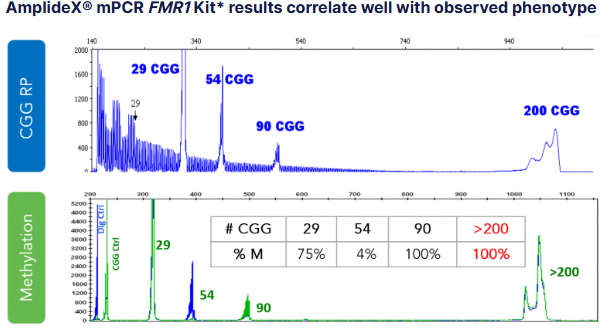
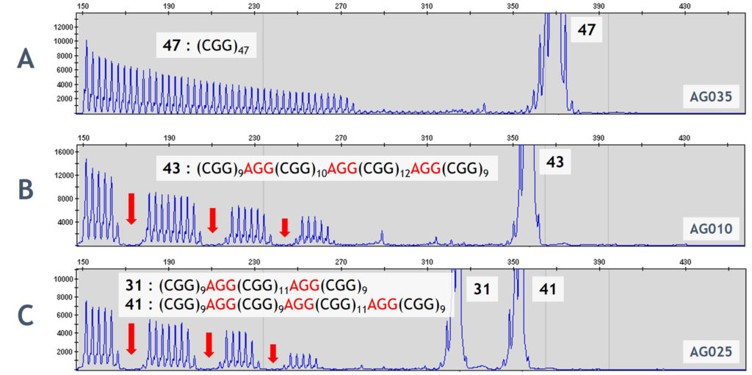
FMR1: CGG repeat quantification and AGG interspersions prevent underestimation of expansion risk.
Clear identification of carriers allows genetic counseling for reproductive planning.
Couples can understand actual reproductive risk, not just probability guesses.
Supports decisions like preimplantation genetic testing (PGT) or prenatal screening.
Reduces need for repeat testing due to inconclusive results.
One comprehensive test per lifetime is enough for most patients.
Reduces lab and patient burden while maintaining clinical confidence.
SMA: SMN2 copy number and c.859G>C modifier are reported, allowing better prediction of severity in offspring.
CFTR: Recognizes CF-related conditions like CBAVD and pancreatitis.
FMR1: Anticipation and stability assessed via AGG interspersions.
Testing Challenges:
Multiple benign variants exist; standard tests can report irrelevant findings.
Accurate carrier detection is critical for reproductive counseling.
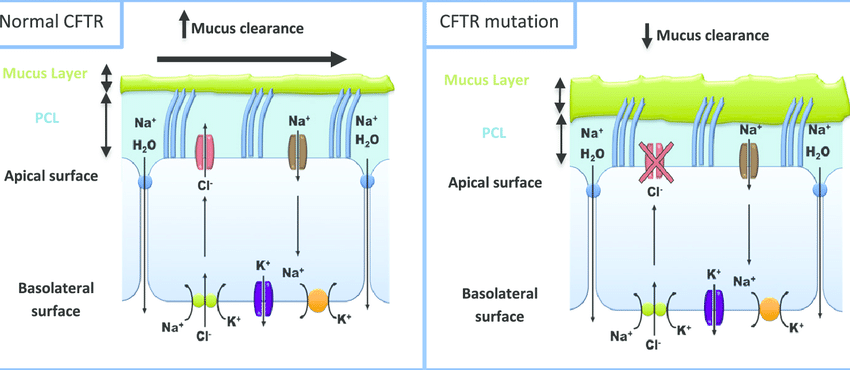
Targeted pathogenic variant testing: Detects only known CF-causing variants, ensuring clinically relevant results.
Provides reproductive risk assessment for couples planning pregnancy.
Testing Challenges:
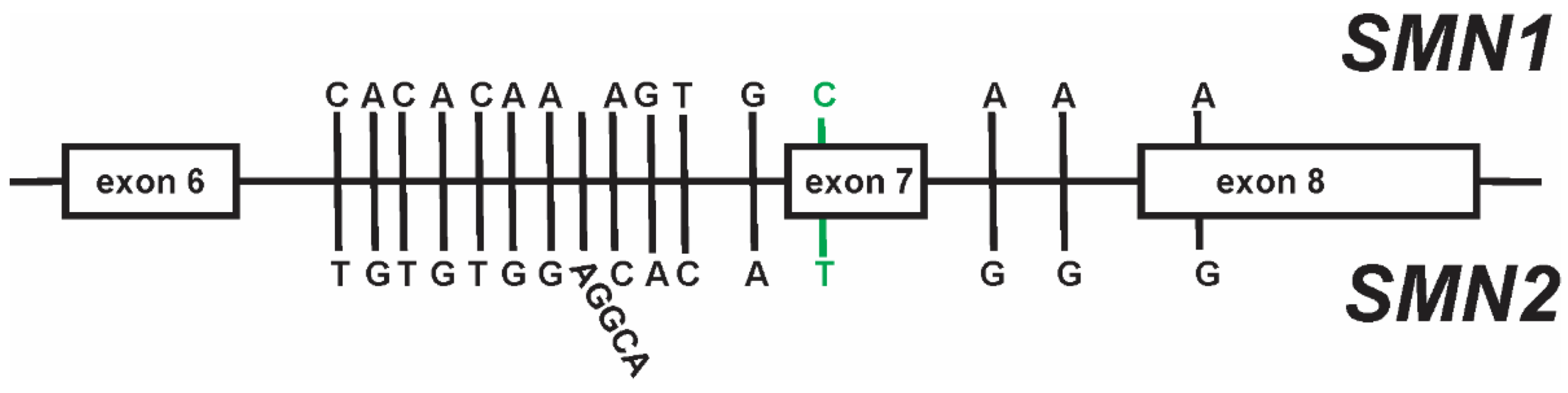
Short-read NGS cannot reliably distinguish SMN1 vs SMN2 due to 99% sequence homology.
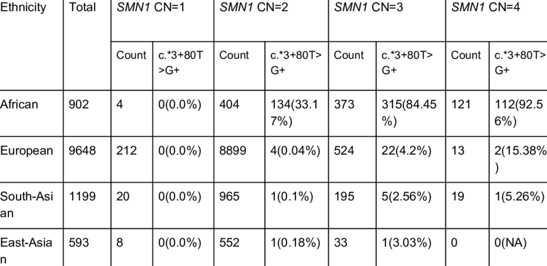
Predicting 2+0 carriers (cis duplication) is complex and ethnicity-dependent.

qPCR for Copy Number Analysis
This method can determine whether a patient has one or two copies of SMN1 on a chromosome, which is critical for identifying carriers. However, qPCR alone cannot distinguish cis from trans configurations, so additional analysis is required for complex carriers.
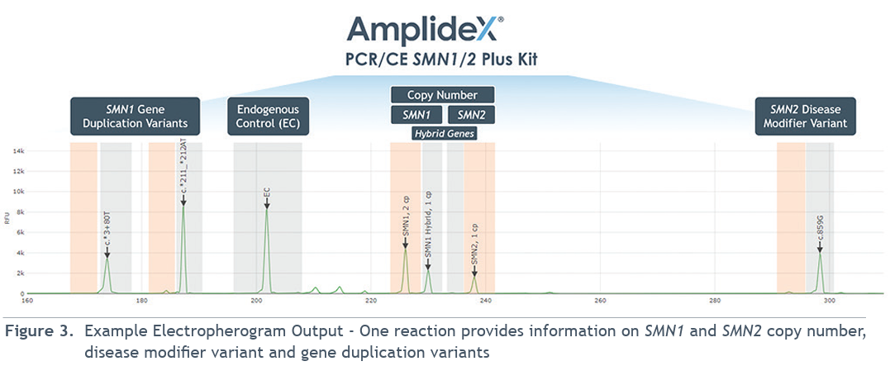
Franklin’s Paralogous Sequence Variant (PSV) bioinformatic approach
By leveraging at least 15 PSVs that differentiate SMN1 from SMN2, reads can be confidently mapped to the correct gene. This enables accurate detection of carriers, even in regions of high sequence homology that typically confound standard NGS.

Clinical Genomics Manager - ANZ & Country Manager - NZ
Contact me today to request a test, submit samples, or get more information — and provide your patients with accurate, actionable reproductive risk insights in a single, reliable report.
“Validated, reliable, and designed for clinicians who need results they can trust.”
For 2+0 carrier prediction, haplotype analysis is performed using SNPs known to be associated with duplicated cis SMN1 copies. This method is ethnicity-aware, improving predictive accuracy across diverse populations. Combined with qPCR and PSV mapping, it provides a highly reliable assessment of carrier status.
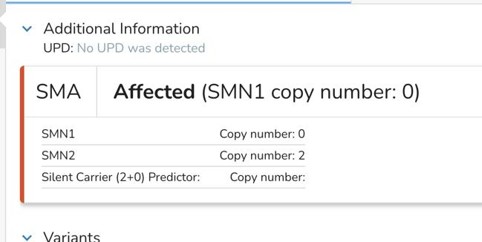
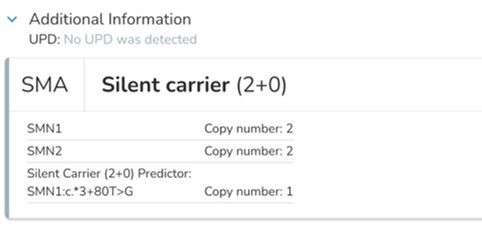
Testing Challenges:
Short-read NGS cannot reliably quantify CGG repeats or assess AGG interspersions.
Inaccurate measurements can misestimate reproductive risk.
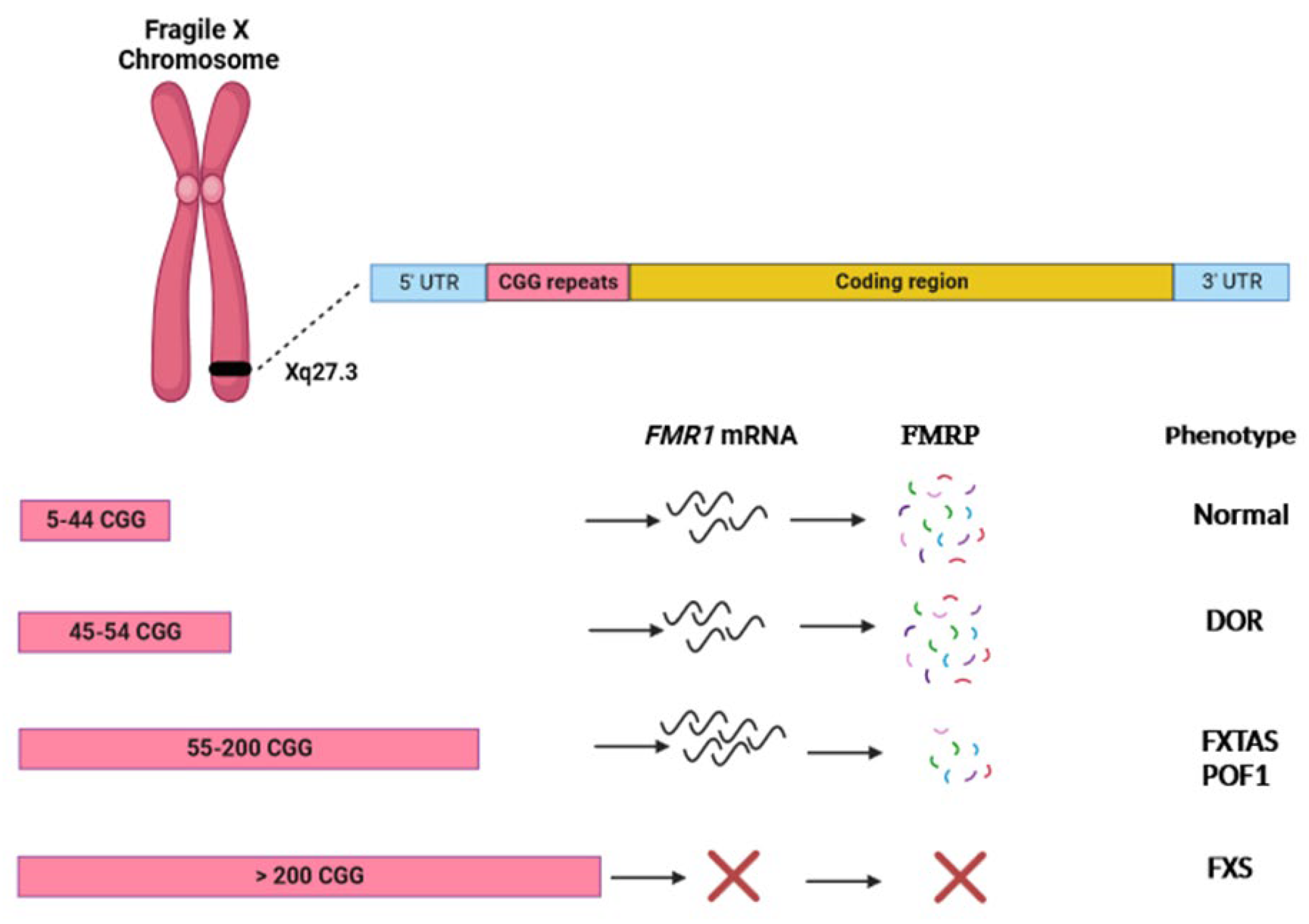
PCR combined with Southern blot analysis. PCR allows precise measurement of CGG repeat length, while Southern blot can detect very large expansions and assess methylation status if needed. Some advanced assays also include AGG interspersion analysis, enabling clinicians to evaluate repeat stability across generations and provide more nuanced reproductive counseling.
By integrating PCR, Southern blot, and optional AGG analysis, this approach provides accurate carrier identification, precise repeat quantification, and predictive insights into disease anticipation. It effectively addresses the mapping and repeat-sizing limitations of NGS, allowing clinicians to counsel families with confidence and make informed reproductive decisions.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.
Simply add your details below and get quick access to the brochure.